August 22, 2023 report
This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Mitochondrial DNA study offers several new findings, reveals confounding factor in previous research
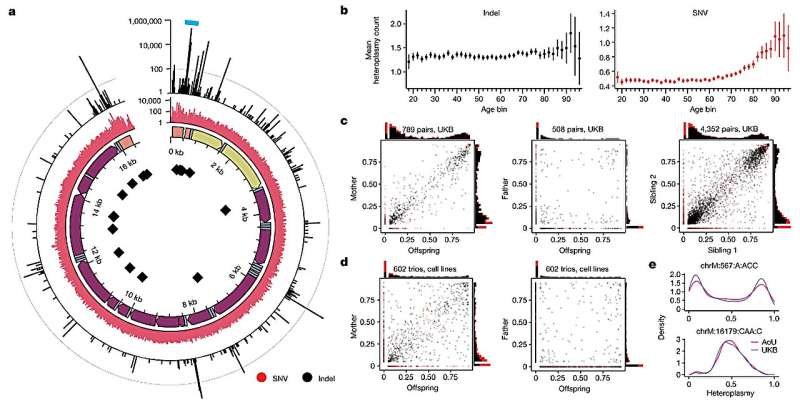
Researchers at the Broad Institute of MIT and Harvard, Cambridge, have interrogated several interconnected questions related to mitochondrial DNA (mtDNA) heteroplasmy and the influence of nuclear genetics.
In their paper, "Nuclear genetic control of mtDNA copy number and heteroplasmy in humans," published in Nature, the team provides a comprehensive understanding of the molecular mechanisms behind mtDNA interactions and their potential implications for human health and evolution.
Mitochondria are cellular organelles that house their own DNA, with thousands of copies of mtDNA in each cell. Each mtDNA molecule is built from 16,569 nucleotides—compared to the ~3 billion nucleotides needed for each copy of the nuclear genome (nuDNA). The mtDNA is inherited maternally and encodes 13 essential cellular energy production proteins, ribosomal RNAs, and transfer RNAs required for mitochondria function.
Heteroplasmy
The study investigates the role of heteroplasmy, in which only some of the mtDNA copies in a cell carry a mutation creating a mixture of different mtDNA alleles within an individual. Some mutations can be a disease risk factor and have been associated with premature aging.
The study finds that individuals carrying harmful mtDNA mutations at low heteroplasmy levels displayed intermediate disease traits. This suggests that the level of heteroplasmy can affect the manifestation level of a disease.
The researchers also find that mutations inherited or acquired with age manifest differently. Insertions and deletions of mtDNA tended to be inherited maternally and shared across siblings, while single-nucleotide variants predominantly accumulate somatically with age.
Influence of nuclear genetics
The study explored how common genetic variations in the larger nuclear genome might affect mtDNA copy number (mtCN), a measure of the number of mitochondrial genomes per cell. MtCN decreases with age, and this decline has been associated with specific nuclear genetic variants.
Nuclear genetic variants were found to influence the levels of mtDNA indels, possibly affecting the replicative efficiency of mtDNA molecules with different alleles. The levels of mtDNA in cells have previously been associated with diseases.
Confounding composition
Unexpectedly, the study found that approximately 25% of the variation in mtCN between individuals can be attributed to differences in blood-cell composition. This suggests that the composition of different types of blood cells may be affecting mtCN levels in individuals.
Previous studies have reported associations between low blood mitochondrial DNA copy number and common diseases, including type 2 diabetes, myocardial infarction, stroke, hypertension, and dementia. After adjusting for blood-cell composition and other covariates, the associations between mtCN and certain diseases could not be found in the UK Biobank cohort.
Intrigued by this finding, the team extended the analyses to 24 more common diseases, finding that, in total, 20 initially showed significantly increased risk with reduced mtCN. After correction for blood cell composition, the correlations disappeared for all traits except osteoarthritis.
Associations with four cardiovascular disease traits flipped their risk correlation.
If the previous associations between low mtCN and diseases are a secondary downstream effect tied to blood cell composition, it would explain how previous researchers may have been confounded. This is a critical finding, as eliminating causal disease associations can also eliminate future research that follows an errant path of looking for disease mechanisms in the wrong location.
The study also illustrates the revealing nature of large and comprehensive research samples. It is unclear if the findings of the study would have been confidently revealed in a study of a few hundred or even a few thousand individuals. UKBiobank and AllofUs provided the researchers with mtCN and heteroplasmy data across approximately 300,000 individuals spanning six ancestry groups.
More information: Rahul Gupta et al, Nuclear genetic control of mtDNA copy number and heteroplasmy in humans, Nature (2023). DOI: 10.1038/s41586-023-06426-5
© 2023 Science X Network