This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Discovery of protein orientation helps scientists understand Parkinson's disease
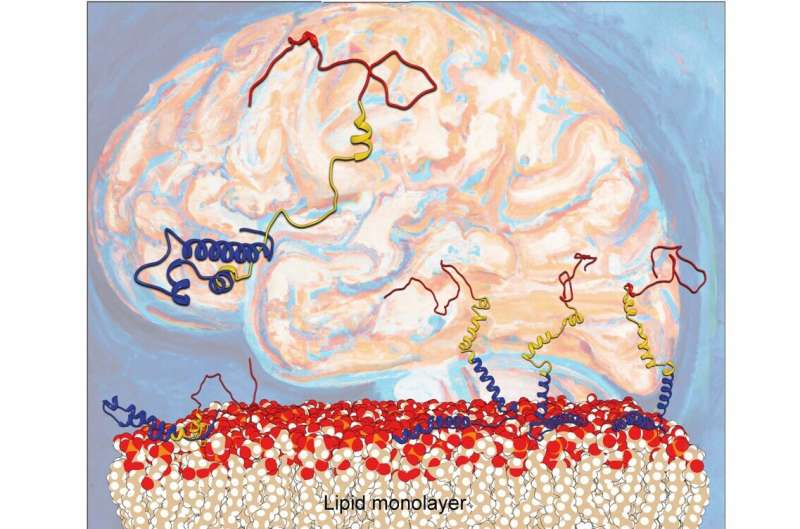
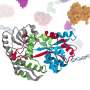
For several years, scientists have known that Parkinson's disease is related to misfolding of the protein alpha-synuclein. Deposited aggregates of a protein called alpha-synuclein (α-syn) can damage and kill nerve cells, leading to neurological dysfunction. It has been known that lipid layers on cell surfaces can accelerate the misfolding process; however, the microscopic mechanisms involved have been a mystery.
Now, scientists from the Departments of Chemistry and iNANO at Aarhus University have unraveled what actually happens when alpha-synuclein binds to lipid layers. A research team headed by Steven Roeters and Tobias Weidner shows how the lipid surface influences the orientation and folding of α-syn.
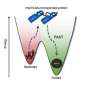
The Aarhus researchers report in Nature Communications that the orientation of α-syn changes at elevated protein concentrations, going from the previously reported flat geometry to a hitherto unknown upright conformation. As a result, the α-syn molecules can refold into dangerous aggregates more easily.
Understanding α-syn folding is of great importance to medical researchers from many different perspectives. Although neurodegenerative diseases are a rapidly growing problem in our aging societies, they still cannot be cured. The refolding of proteins and their interactions with cell surfaces is a much-debated topic in current Parkinson's research.
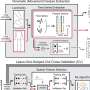
To understand how α-syn behaves at surfaces, the researchers produced and used synthetic proteins, placing them in a single layer on a model nerve cell surface. To investigate the binding, motions and folding of this nanometer-thin layer of proteins, the team used a laser-based spectroscopic method called sum frequency generation spectroscopy. The complex datasets were disentangled using a computational method specifically developed with the department's computer simulations group.
Thanks to these new findings, it may now be possible to follow the misfolding process and the working mechanisms of potential medical drugs in molecular detail. "In the future we might use this method to screen for effective therapies to prevent the disease," says Steven Roeters, who is now a research associate at Amsterdam UMC.
Encouraged by the positive results, the Aarhus research groups want to extend their investigations to other surfaces which α-syn might encounter and which reflect the way our society affects our surroundings, e.g. plastic contaminants.
"We plan to examine the interactions of α-syn with artificial materials, such as microplastics to understand the impact of environmental conditions on Parkinson's," explains Associate Professor Tobias Weidner, whose working group at Aarhus University specializes in the characterization of proteins at surface and nanomaterials.
More information: Steven J. Roeters et al, Elevated concentrations cause upright alpha-synuclein conformation at lipid interfaces, Nature Communications (2023). DOI: 10.1038/s41467-023-39843-1