This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Unveiling asthma's molecular secrets: How blood molecules influence airway processes
A new study by researchers at the Icahn School of Medicine at Mount Sinai has unraveled the intricate molecular interplay between systemic processes within the blood and localized processes within the airways of individuals with asthma.
This pioneering research opens doors to potential novel treatments targeting specific molecules, with the aim of providing more effective relief for asthma patients. The research findings were published today in Genome Medicine.
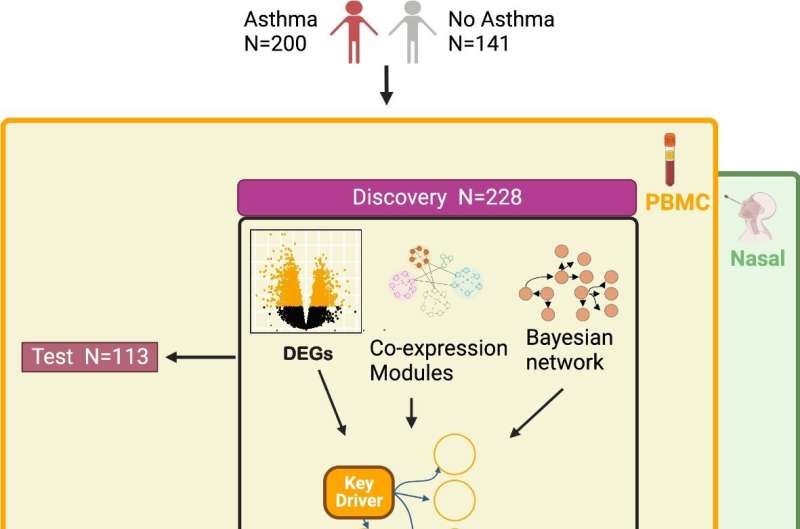
The study, which examined 341 participants comprising individuals with persistent asthma and non-asthmatic controls, used advanced transcriptomic sequencing techniques to analyze blood and nasal samples. Through this comprehensive approach, the researchers unveiled crucial molecules and processes that hold the key to understanding asthma better. Within the blood, the NK cell granule protein and perforin emerged as central players. In the nasal passages, the G3BP stress granule assembly factor 1 and InaD-like protein took on pivotal roles.
Notably, the study underscored the profound influence of blood molecules on asthma by virtue of their effects on nasal molecules.
Supinda Bunyavanich, MD, MPH, MPhil, the Mount Sinai Professor in Allergy and Systems Biology, Deputy Director of the Elliot and Roslyn Jaffe Food Allergy Institute, and a senior author of the study, highlighted the importance of this holistic approach: "Our findings represent a groundbreaking connection between systemic factors in the bloodstream and localized factors in the airways, working collaboratively to drive asthma. This discovery marks a significant stride towards understanding core mechanisms of asthma, transcending the conventional focus on only the airways."
However, Dr. Bunyavanich also sounded a note of caution, saying, "While these revelations provide invaluable insights into the molecular framework of asthma, it's imperative to acknowledge that further research is needed before these breakthroughs can translate into immediate therapies."
Key highlights:
- The study offers an integrated perspective on the intricate relationship between systemic processes and airway-specific mechanisms in asthma.
- Prominent blood molecules, including the NK cell granule protein and perforin, appear to exert their influence on asthma through their interactions with nasal molecules like the G3BP stress granule assembly factor 1.
- The findings chart a path for future research endeavors, guiding the development of targeted asthma therapies that modulate these specific molecules.
- Given that asthma affects millions globally, the implications of this research are far-reaching. This study paves the way for a deeper comprehension of the disease, instilling optimism for the emergence of more efficacious therapeutic strategies in the foreseeable future.
More information: Lingdi Zhang et al, Integrated study of systemic and local airway transcriptomes in asthma reveals causal mediation of systemic effects by airway key drivers, Genome Medicine (2023). DOI: 10.1186/s13073-023-01222-2





















