This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Army of specialized T cells may trigger asthma attacks in older men
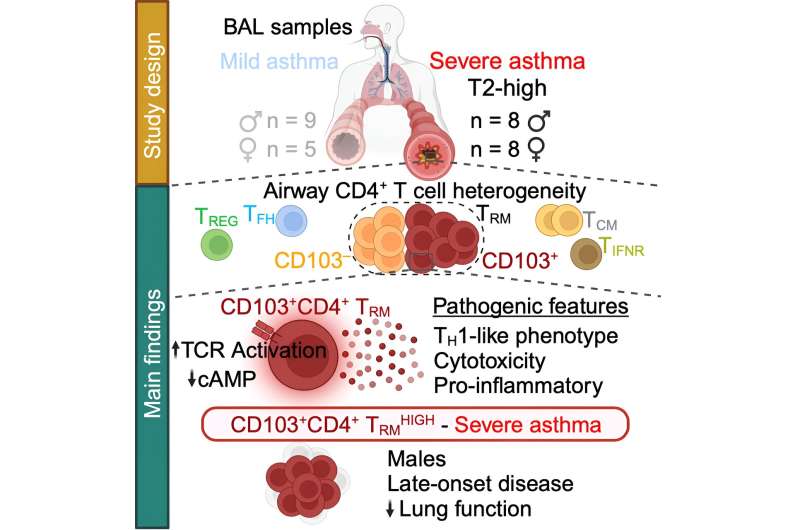
Scientists from La Jolla Institute for Immunology (LJI) and The University of Southampton, UK, have uncovered a group of immune cells that may drive severe asthma. These cells, called cytotoxic CD4+ tissue-resident memory T cells, gather in the lungs and appear to possess the molecular weaponry to cause the most harm in men who developed asthma later in life.
"If you are male and you develop asthma after age 40, there's a high chance this T cell population is in your lungs," says LJI Research Assistant Professor Gregory Seumois, Ph.D., who co-led the study with LJI Professor Pandurangan Vijayanand, M.D., Ph.D.
The scientists uncovered these pathogenic T cells thanks to volunteers enrolled in the NHS-clinic based WATCH study, which follows hundreds of asthma patients of different ages, sexes, and disease severities. By following patients over many years, and analyzing their immune cell populations, researchers are making new connections between asthma symptoms and immune cell activity.
"Once you understand the role of cells like these T cells better, you can start to develop treatments that target those cells," says WATCH study director Ramesh Kurukulaaratchy, BM, DM, FRCP, Associate Professor at the University of Southampton and researcher at the NIHR Southampton Biomedical Research Centre.
The scientists now hope to learn more about these cells—and their role in asthma development—as they work to bring personalized therapies to asthma patients.
How harmful T cells drive asthma
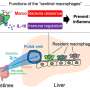
So how might these harmful, or "cytotoxic," CD4+ tissue-resident memory T cells trigger asthma in older men? The problem may come down to a combination of immune cell "memory" and an absence of helpful cells in the lungs.
These T cells are called "memory" cells because they react to molecules that the body has previously fought off. This kind of immune cell memory helps protect the body from viruses and bacteria. Unfortunately, the same T cell memory is a big problem for asthma patients. Their misguided T cells see harmless molecules, such as pollen, and mount a dangerous inflammatory response.
The new research suggests asthma patients with these T cells in their lungs may be primed for hard-to-treat, and potentially fatal, asthma attacks.
The scientists aren't sure why these T cells tend to cause problems for older men. Seumois is interested in solving this mystery through LJI's Center for Sex-Based Differences in the Immune System. "We know that females have a different immune landscape," says Seumois. "So we're interested in investigating this question further."
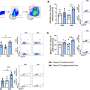
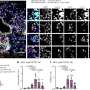
LJI graduate student Sara Herrera de la Mata used a technique called single-cell RNA sequencing to learn more about immune cells in these patients. She found that certain anti-inflammatory T cells that should counteract severe asthma symptoms were scarce in airway fluid samples (BAL samples) from this patient group.
Instead, men who developed asthma later in life had an overwhelming number of potentially harmful T cells. Their lungs should have been home to a diverse bunch of CD4+ T cell types, and yet for this group, more than 65 percent of their cells were cytotoxic CD4+ tissue-resident memory T cells.
"Normally, a doctor would give a severe asthma patient a steroid or biologics therapy to dampen their immune response, and that should be it," says Herrera de la Mata, who served as co-first author of the study. "But these cells may not respond to these treatments."
Spotting this immune cell imbalance was an important clue that this patient group represented a new asthma subtype.
Discovery could lead to personalized asthma treatments
The sequencing work at LJI gives scientists a "biomarker" to help them detect cytotoxic CD4+ tissue-resident memory T cells in more patients going forward.
In fact, finding this biomarker represents a "paradigm shift" in asthma research, says Kurukulaaratchy. Before now, scientists and clinicians separated asthma patients into just two groups: "T2 high" and "T2 low." This dogma isn't helpful for patients or clinicians, says Kurukulaaratchy. As a clinician, he knows that the T2 high group actually includes a huge range of patient demographics and symptoms.
"We've studied a subgroup of male patients who developed asthma later in life—and they do badly. They need a lot of treatments, such as high doses of toxic steroid treatments," says Kurukulaaratchy.
In a study published earlier this year, the research team showed the importance of "drilling down" to identify many more asthma patient subgroups. Their analysis reveals that 93 percent of WATCH subjects with severe asthma were in the T2 high category. Clearly, T2 high is a broad category.
"T2 high" is too broad, in fact, to really help doctors narrow down treatment strategies for individual patients, explains study co-author S. Hasan Arshad, MBBS, DM, FRCP, Chair in Allergy and Clinical Immunology at the University of Southampton, researcher at the NIHR Southampton Biomedical Research Centre, and Director of The David Hide Asthma and Allergy Research Centre, Isle of Wight.
"We have to think of severe asthma as having different subtypes, and the treatment has to be tailored according to these subtypes—because one size does not fit all," says Arshad.
The researchers' mission now is to use sequencing tools and other techniques to discover additional biomarkers and asthma patient subtypes. Seumois says he is looking forward to continuing his collaboration with Southampton scientists and the WATCH cohort. He's also making plans to examine immune cells in more patient groups, including a cohort that includes African American and Hispanic patients, two understudied demographic groups known to be at a higher risk for developing severe asthma.
"By looking at clinical features of asthma and biometrics, we're finding things that have never been shown before," says Seumois.
The research is published in the journal Med.
More information: Sara Herrera-De La Mata et al, Cytotoxic CD4+ tissue-resident memory T cells are associated with asthma severity, Med (2023). DOI: 10.1016/j.medj.2023.09.003