This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
trusted source
proofread
Artificial intelligence aids fight against acute myeloid leukemia

When Mauricio Ferrato completed his doctorate in computer and information sciences at the University of Delaware a few months ago, he made his mark in more ways than one.
Ferrato played a pivotal role in a research collaboration involving UD and Nemours Children's Health that used artificial intelligence to home in on the most effective drug therapies for patients with acute myeloid leukemia (AML), an aggressive blood cancer.
The work, which was published earlier this year in the journal Bioinformatics Advances, is another step forward in the drive toward precision medicine, where treatment will be personalized to a patient's unique genetic profile, with greater effectiveness and fewer adverse impacts.
According to the Leukemia and Lymphoma Society, about 20,000 new cases of AML emerge each year in the United States, and more than 11,000 people die from the disease annually.
It affects both children and adults and occurs when the body makes too many immature blood cells, called myeloid blasts, that can't develop into normal white blood cells.
These abnormal cells grow out of control and crowd out healthy cells in bone marrow. From there, they can spread to the lymph nodes, brain and other organs, causing a broad range of symptoms, from fatigue and shortness of breath, to joint pain, frequent infections and weight loss.
The blood cancer progresses rapidly, so early diagnosis is critical. The five-year survival rate for patients after diagnosis is 31.7%, according to the National Cancer Institute.
Using genetic data from 451 patients made available through the BeatAML initiative, Ferrato used machine learning, a form of artificial intelligence, to help determine if a person with AML will be a "high responder" or a "low responder" to any of 100 different drug therapies. Then the team was able to "reverse engineer" the findings and track the pathways back to a particular gene in a patient and determine if that gene was creating a protein that impacts the cancer or a protein that resists a specific drug.
This foundational research may help lay the groundwork for more promising outcomes for patients. The researchers also hope to explore the impact of their strategy on other types of cancer datasets and drug therapies.
Putting machine learning to the test
Machine learning operates on algorithms—sets of instructions that allow computers to make predictions and decisions based on data—without being explicitly programmed to do so. These algorithms help identify patterns and relationships from massive amounts of data and generate computer models of the findings.
This field of artificial intelligence was critical to the AML project, which was co-led by Ferrato's doctoral adviser, Sunita Chandrasekaran, associate professor of computer and information sciences at UD, and Erin Crowgey, previously director of medical bioinformatics at Nemours Children's Health, and currently associate director of bioinformatics at Incyte, a biopharmaceutical company headquartered in Wilmington, Delaware. Adam Marsh, associate professor in UD's School of Marine Science and Policy, also was involved, along with colleagues from Emory University and the University of California San Diego.
While Ferrato brought plenty of machine learning muscle to the project, he wasn't always drawn to computer science.
Originally from Venezuela, Ferrato came to Delaware when his parents moved to the state when he was 12 years old.
"UD was the best option for me—it allowed me to live close to family, the research has a strong reputation and the campus is beautiful," he said. "I actually wanted to go into sports journalism when I started, but then I ended up working with Sunita as an undergrad, mostly in high-performance computing."
Chandrasekaran had a big impact on Ferrato, and he stayed on for his master's and doctoral degrees in UD's Department of Computer and Information Sciences.
Ferrato got involved in the AML project when Crowgey, a UD doctoral alumna in bioinformatics, was working at Nemours Children's Health and had received funding from the Lisa Dean Moseley Foundation to pursue research on pediatric patients with the disease.
"We had funding through the grant to bring aboard a Ph.D. student, and Mauricio was a perfect fit," Crowgey said. "Our goal was to answer the question: Could you predict before treatment that a person would respond to a given drug?"
Crowgey compared the work to having a lot of marbles in a jar and figuring out which marble is the most important.
"That's what feature selection is about in machine learning," she said. "Once you find that marble, it may be large or oblong. How will it roll? It's a way to take a lot of data that an individual can't easily interpret and create an algorithm to pull out what's meaningful from 20,000 genes in the genome, in this case, and show how a person with AML will respond to treatment."
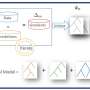
Ferrato used SHAP (short for SHapley Additive exPlanations), a tool used in game theory, to map a particular feature back to its biological equivalent. So SHAP would select the top 30 features, each representing a gene, and then pathway analysis would show what that gene was affecting, such as creating a protein that resists an anti-cancer drug.
He put in many hours writing computer code in Python and running models on UD's DARWIN high-performance computer.
"We looked at six different models for 100 different drugs, and then we'd have to run the models multiple times to validate them, checking to see if the results were consistent. We had to run 3,000 to 4,000 models to generate results, with each model taking about an hour to run," he explained.
The promise of AI
As a computer scientist, Ferrato said he didn't know all the biological terms associated with the project, such as transcriptomes, gene expression, RNA and the background on AML.
Crowgey mentored him. In turn, he helped her better understand machine learning.
Soon after he completed his doctorate, Ferrato began working for NVIDIA as a solutions architect. He had interned there during graduate school, using his computer science skills to find the optimal way to site wind turbines in a wind farm to generate the most energy possible.
"I like work that is applied to a real-life problem that helps humanity in some way," he said.
Working together, researchers from multiple disciplines can solve large-scale problems using artificial intelligence that they just couldn't tackle before. Team science is a key, Crowgey said.
And as far as future applications for artificial intelligence, the sky's the limit.
"We need to be really smart about how we develop and implement these applications," Crowgey said. "We all use AI every day, but we just don't think of it that way. Your cellphone has all kinds of cool AI on it. This work for AML patients is powerful and impactful."
Chandrasekaran also is a staunch advocate of interdisciplinary problem-solving, working with industry and academic partners. It's a hallmark of the new AI Center of Excellence (AICoE), which she now co-directs at UD.
"Working with our collaborators, Mauricio and I learned a lot about the impact machine learning can have in precision medicine. The outcomes were fascinating," Chandrasekaran said.
"The explosive growth we see in generative AI tools underscores the need to ensure that our next-generation workforce is prepared to use these tools," she noted. "To that end, our AI Center of Excellence at UD, which works with investigators across various disciplines to provide AI solutions, also recently launched a Graduate Certificate in AI. It is open to UD students, as well as to professionals outside UD."
Now that the AML research results have been published, what happens next?
"This work is laying the foundation, the infrastructure, the technology for the future," Crowgey said. "It will take the community as a whole, bringing together academia, hospitals and the biopharma industry, to drive precision medicine forward."
More information: Mauricio H Ferrato et al, Machine learning classifier approaches for predicting response to RTK-type-III inhibitors demonstrate high accuracy using transcriptomic signatures and ex vivo data, Bioinformatics Advances (2023). DOI: 10.1093/bioadv/vbad034