This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Reverse metabolomics: New method finds biomarker for inflammatory bowel disease
In recent years, microbiome research has started to shift its focus from the microbes themselves to the molecules they produce. After all, it's these molecules that directly interact with human cells to influence a person's health. However, trying to identify which molecules are being made by a person's microbiome is quite challenging. A typical metabolomics study can only characterize about 10% of the molecular data from a human microbiome sample.
In a new study published in Nature, microbiome experts at University of California San Diego debut a new approach they call "reverse metabolomics." The technique combines organic synthesis, data science, and mass spectrometry to better understand what molecules are being secreted by the microbiome and how they affect human health.
In their first application of reverse metabolomics, the scientists found hundreds of molecules that had never been observed in the human body before. Using this novel data, they were able to identify a new metabolomic signature for inflammatory bowel disease (IBD). The authors say these molecules could someday serve as a biomarker for diagnosing IBD or as a potential therapeutic target to help treat the disease.
"We know the microbiome is important, but we don't know what kinds of molecules the microbes produce or how influential they are in the human body," said senior author Pieter C. Dorrestein, Ph.D., professor at Skaggs School of Pharmacy and Pharmaceutical Sciences at UC San Diego. "Reverse metabolomics helps us evaluate whether specific molecules can be found in the samples, predict which microbes are producing them, and relate these metabolomic signatures to health and disease."
In a typical metabolomics study, researchers will use a tool called mass spectrometry to look for different molecules in a sample. In this technique, each molecule has its own "barcode" that it can be identified by. Still, scientists need to know what these barcodes represent to describe the contents of a sample, which remains a challenge.
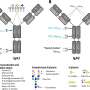
In the new study, researchers in the Dorrestein Lab took a backward approach. First author Emily C. Gentry, Ph.D., now an assistant professor at Virginia Tech, used organic synthesis to first produce thousands of different synthetic molecules from four classes of interest and then defined each of their barcodes.
The researchers then utilized publicly available metabolomics data, including the ones previously collected through the Crohn's & Colitis Foundation, and searched for the new barcodes in that data. The findings revealed 145 of the synthesized bile acids were present in biological samples from public data, 139 of which had never been described before.
"If you read a biology textbook, none of these molecules will be in it," said Dorrestein. "Not only are they new to our understanding of human physiology, they're entirely new to science, which is pretty amazing."
Gentry and colleagues then compared the metabolomic signatures of samples from different patient populations and found a strong association between a synthesized class of microbial molecules—bile amidates—and IBD. This association was then validated across multiple cohorts, supporting the idea that these molecules are likely involved in the pathology of IBD.
Looking closer, the scientists noticed that certain bile amidates were elevated in patients with Crohn's disease, specifically when they had active symptoms, but this was not the case for patients with ulcerative colitis. Patterns like these could one day be used to help differentiate and diagnose specific types of IBD.
The researchers then began to explore how these molecules might be influencing gut health. Additional experiments showed that several bile amidate compounds may promote gut inflammation by dysregulating T cell function. For example, one microbial compound produced a six-fold increase of a key cytokine known to be involved in the pathogenesis of Crohn's disease.
"We're using organic synthesis and data science to better understand how our bodies work on a molecular level," said Gentry. "We're also one of the first studies to discover new human molecules using publicly available metabolomics data. As more metabolomics data becomes publicly available, reverse metabolomics will become even more informative."
The authors say the molecules they've described could one day inspire new therapeutics for treating IBD. For example, patients might be treated with pills containing live microbes that secrete specific molecules or drugs that inhibit the enzymes these disease-associated molecules interact with.
"This is a remarkable achievement derived from our precision nutrition initiative, in which Dr. Dorrestein previously demonstrated that reverse metabolomics could identify food metabolites associated with disease severity in patients with IBD," said Andrés Hurtado-Lorenzo, Ph.D., senior vice president of Translational Research & IBD Ventures at the Crohn's & Colitis Foundation. "Now, this groundbreaking work has further progressed to discovering new metabolites that hold potential for both diagnostic and therapeutic applications in IBD."
More information: Emily C. Gentry et al, Reverse metabolomics for the discovery of chemical structures from humans, Nature (2023). DOI: 10.1038/s41586-023-06906-8