This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
proofread
Researchers create comprehensive transcriptional atlas of human adenomyosis
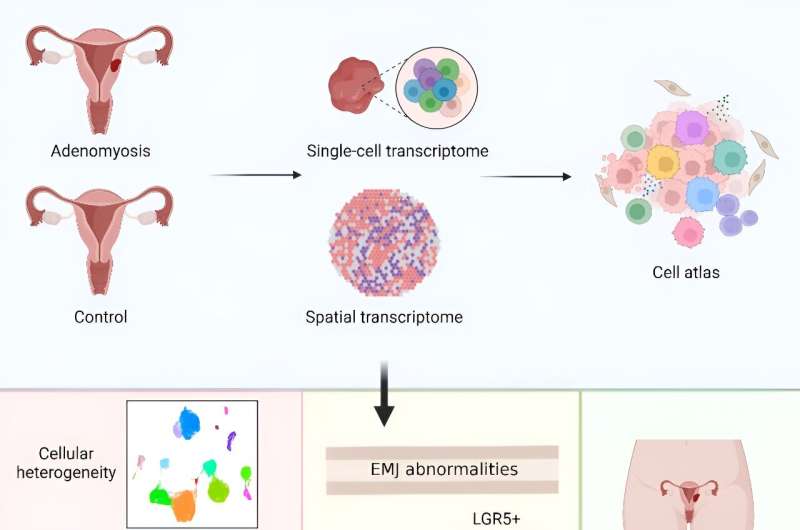
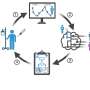
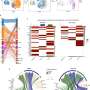
Adenomyosis is a poorly understood gynecological disorder with limited treatment options. A new study employed single-cell RNA sequencing and spatial transcriptomics to map transcriptional alterations across different regions of the uterus in adenomyosis patients and controls.
It highlights unique epithelial and stromal subpopulations and aberrant signaling pathways involved in adenomyosis, offering potential targets for precise diagnostics and therapeutics. The research is published in the journal Protein & Cell.
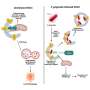
The researchers found that unique epithelial (LGR5+) and stromal (PKIB+) subpopulations were identified in adenomyotic lesions, supporting a complex interplay between "invagination" and "metaplasia" theories. Additionally, WFDC1+ stromal progenitor cells might act as precursors to lesion-specific stromal cells, indicating a sophisticated mechanism of lesion formation.
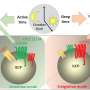
Abnormal angiogenic signaling and endothelial cell heterogeneity were also observed in lesions. Notable pathways involved include vascular endothelial growth factor and angiopoietin, highlighting disrupted angiogenic processes in these tissues.
The team also identified altered cell–cell communication between ectopic and eutopic endometrium, particularly within adenomyotic lesions. These aberrant signaling pathways, including pleiotrophin, TWEAK, and WNT cascades, suggest modified signaling dynamics in the lesions.
This study advances the understanding of adenomyosis by providing a comprehensive single-cell and spatial transcriptomic landscape. It reveals unique cellular subpopulations and signaling aberrations within adenomyotic lesions, supporting both invagination and metaplasia theories.
These insights offer the potential for developing precise diagnostic and therapeutic strategies, emphasizing the need for clinical validation of these findings.
More information: Tao Chen et al, Comprehensive transcriptional atlas of human adenomyosis deciphered by the integration of single-cell RNA-sequencing and spatial transcriptomics, Protein & Cell (2024). DOI: 10.1093/procel/pwae012