Cancer drugs taking shape
In the era of molecular medicine, potentially active compounds for use in cancer therapies can be identified faster than ever before. Yet pinpointing the molecular target of an anticancer compound and deducing its mode of action remains a painstaking process. Yushi Futamura, Hiroyuki Osada and colleagues from the Chemical Biology Department of the RIKEN Advanced Science Institute have now discovered that anticancer compounds induce a shape change in target tumor cells that is directly related to a compound's mode of action.
The cancer cell shape-shifting phenomenon was discovered by Futamura during a study of cells treated with the anticancer drug vinblastine, a tubulin inhibitor. He recalls being shocked by the dramatic morphological shift that the drug induced in the cells. "I remember thinking, 'Are these the same cells?'" he says.
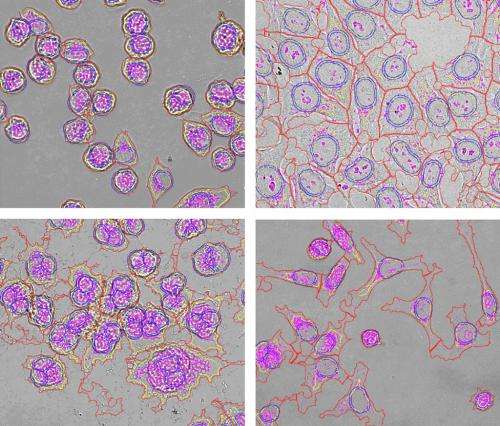
Inspired by his initial observations of cell shape changes, Futamura and his colleagues began testing other known tubulin inhibitors and found that each had the same morphological effect—turning treated cells into flattened disks. Systematically examining other anticancer drug classes, the team discovered that each class induces its own characteristic changes in cell shape (Fig. 1). "We thus came up with the idea of making an encyclopedia of cell morphology, which we call Morphobase," says Futamura.
Morphobase now includes the results of tests on over 200 anticancer compounds with known modes of action, where shape changes in treated cells are quantified against 12 morphological parameters.
The researchers evaluated the potential of Morphobase as a predictive tool for rapid determination of anticancer compound modality by testing three anticancer compounds with unknown modes of action. By comparing the observed changes in treated cells with results in the database, they quickly established that all three were tubulin inhibitors. Further tests using conventional techniques confirmed the result.
Osada, Futamura and their colleagues are continuing to use Morphobase to establish the mode of action of novel anticancer compounds with promise as potential drug leads. The researchers are also refining and expanding the database by broadening the drug reference set and the range of drug doses for which morphology data is maintained. "These improvements and extensions to the system could improve its capability to identify the molecular mechanisms of drug activity, and in discovering novel drug candidates as well," says Futamura. The database has the potential to significantly cut the time it takes to establish the mode of action of novel anticancer compounds, removing a significant bottleneck in drug discovery research.
More information: Futamura, Y. et al. Morphobase, an encyclopedic cell morphology database, and its use for drug target identification. Chemistry & Biology 19, 1620–1630 (2012). dx.doi.org/10.1016/j.chembiol.2012.10.014