Drugs without side effects: Researchers explore novel ways to classify proteins

Janelle Leuthaeuser is on the cutting edge of biophysics. A molecular genetics and genomics Ph.D. student, she is part of a nationwide effort to create a more efficient generation of protein-based drugs.
Proteins are the worker bees of cells. They get rid of waste, transmit cellular signals and carry out the chemical reactions that keep our bodies running. The biomolecules' role as intracellular regulators and their sheer number (there are about 7,000 different types of protein molecules in a typical human body cell) also make them one of the most frequent targets of drugs.
Drugs can help your body fight off disease and infection by speeding up or altering the chemical reactions regulated by proteins. Around half of the drugs currently on the market work by targeting proteins.
Researchers have identified millions of proteins and are discovering new ones faster than ever before. The problem, when it comes to making new and better drugs, is that two proteins can have a similar structure or genetic sequence but react very differently to the same chemical compound. This can lead to adverse drug side effects that range from mildly discomforting to potentially life-threatening. In fact, more than three million serious adverse reactions to prescription drugs occur every year. There are more than 100,000 deaths from these reactions per year, placing prescription drug use as the fourth leading cause of death in the United States, ahead of pulmonary disease, diabetes, AIDS and pneumonia.
"The more information we can get about how specific proteins function the better we can target them with drugs," Leuthaeuser said. "We have the technology to identify a lot of new proteins now, but not to efficiently identify the details of their molecular function."
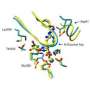
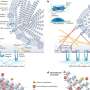
She is working with Jacquelyn Fetrow, Reynolds Professor of Computational Biophysics and Dean of the College, and a national team of researchers to develop an automated program that classifies different proteins by their active site, the location where chemical reactions occur. Contemporary technologies predict protein function based on the full sequence or structure of protein molecules. The Wake Forest led team found this method often vaguely or incorrectly classifies proteins. Leuthaeuser said that proteins can be grouped together by function more accurately when using the sequence and structure directly surrounding the active site rather than the overall structure of a protein molecule.
"When proteins have less than 30-40 percent sequence identity, it is more helpful to look at just the active sites," Leuthaeuser said. "We can more accurately predict their function when we are looking at this small area."
A simple example of how the active site information can be applied in developing more efficient drugs is with aspirin. Leuthaeuser said aspirin affects two similar proteins – COX 1 and COX 2 – that do two very different things. COX 2 is involved with general pain and inflammation while COX 1 regulates swelling in the gastrointestinal tract. Aspirin inhibits both COX 1 and COX 2.
"This is why taking aspirin can give you an upset stomach," Leuthaeuser said. "We would like to inhibit just COX 2 and not COX 1. "There are differences in the active sites of COX 1 and COX 2 that allow drugs to be targeted to COX 2 selectively, decreasing the potential gastrointestinal side effects."
Leuthaeuser is in the process of characterizing active site information for a group of structurally diverse proteins using a computer program developed by members of the Fetrow Lab. While the project is in an early testing and validation stage, she said the eventual hope is that utilizing active site profiling technology will enable accurate and efficient protein function prediction. This could in turn assist pharmaceutical researchers in designing drugs with fewer adverse effects.
Patsy Babbitt, a professor of Pharmaceutical Chemistry at the University of California, San Francisco, has been working with Fetrow on models to predict protein function for five years and is a collaborator on the project. She said automated systems like this will not only make it easier to develop drugs with fewer side effects but will greatly speed up the process of drug development in general. Until recently, Babbitt said the only way to efficiently and accurately identify protein function at the necessary level of molecular detail was to examine them one by one in the lab, a painstakingly slow process that requires a significant amount of time for each protein and costs a significant amount of money.
She said the goal is to bring the time down from a few years to a few weeks.
"The precision will not be as good as individual experiments," Babbitt, said. "But, it will take much less time and money to build an infrastructure to help other scientists form hypotheses and identify the details of protein functional sites."