Different regions, different genetic risk factors

A genetic study of Chinese patients reveals a prevalent risk factor for certain blood cancers not detected in European patients.
The genetic root of cancer often resides in the combined effects of gene mutations that individually exert only a modest effect. A genome-wide association study (GWAS) is one way to learn more about such mutations. Researchers conduct these studies to comb the genomes of large numbers of individuals and identify sequence changes that are meaningfully over-represented in cancer patients relative to unaffected counterparts.
A GWAS by a multinational team of researchers including Jianjun Liu of the A*STAR Genome Institute of Singapore has now revealed a genetic variation that markedly elevates the risk of blood cancer in Chinese populations1. A critical limitation of GWASs conducted to date has been their historic tendency to focus on patients of European ancestry, potentially overlooking important risk factors that may be more prevalent in other ethnic groups. "GWASs in Chinese, South Korean and Japanese populations have uncovered genetic risk factors for nasopharyngeal cancer, gastric cancer and liver cancer that are a lot more common in Asian people," says Liu.
Accordingly, Liu's team partnered with colleagues in Singapore and China to search for genetic variants linked with a subset of blood cancers classified as B cell non-Hodgkin lymphoma (NHL) in East Asian patients. Their work was designed to follow up on a successful search within European patients.
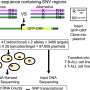
In the initial analysis, the researchers examined 274 Chinese NHL patients and 1,500 healthy controls from Singapore, looking for small genome sequence variations known as single nucleotide polymorphisms (SNPs) that might elevate disease risk. They identified 59 candidate SNPs in this first round, but further rounds of validation in additional cancer patients and a control cohort eliminated all but one of them.
The variant exhibited an extremely robust statistical association with NHL. "It confers a 50 per cent increased risk of developing NHL in individuals that carry this DNA variant as compared to those who do not," says Liu. The SNP resides in the region between two genes, BCL6 and LPP, which have both previously been linked to cancer. BCL6 in particular is known to act within a number of immune cell-specific signaling pathways. Even though the newly identified variant does not alter the genetic sequence, it may act on sequences that critically regulate BCL6 expression.
Liu hopes to further validate this link by searching for BCL6 variants that may be over-represented in European NHL patients, while continuing to delve deeper into the genomes of Asian patients. "We are especially interested in lymphoma subtypes that are more common in Asian populations," he says.
More information: Tan, D. E. K., Foo, J. N., Bei, J.-X., Chang, J. Peng, R. et al. Genome-wide association study of B cell non-Hodgkin lymphoma identifies 3q27 as a susceptibility locus in the Chinese population. Nature Genetics 45, 804–807 (2013). dx.doi.org/10.1038/ng.2666