Researchers reveal the secret of radiation vulnerability
The discovery can help both in predicting the consequences of irradiation and understanding the fundamental patterns of morphogenesis. The work of the researchers has been published in PLoS One journal and is available online.
The scientists - Boris Kuzin, Ekaterina Nikitina, Roman Cherezov, Julia Vorontsova, Mikhail Slezinger, Olga Zatsepina, Olga Simonova, Grigori Enikolopov and Elena Savvateeva-Popova - studied Drosophila flies, in whose genome weak mutations of two different genes were combined. They concluded that these mutations synergistically strengthen their mutual phenotypic expression. In other words, the aggregate effect of these mutations is much greater than that which can be produced by one of them individually.
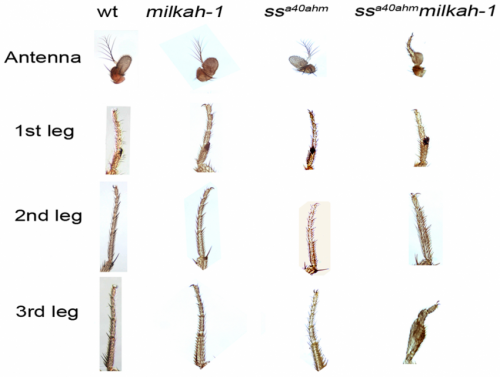
The mutant flies bred by the scientists have a number of significant peculiarities. The experiments have shown that even low doses of X-ray irradiation (not exceeding 10 R) can cause serious defects in those flies' legs.
In contrast, in the flies with normal (unchanged) genome such defects could not be caused even by doses of irradiation hundreds of times higher. What is more, the combination of the two mutations worsened the long-term memory impairment, earlier observed in the flies with only one of the mutations.
In addition, the mutant flies' cells are less resistant to the so-called superoxide radicals.
Superoxide radicals are ions which appear in cells under both normal and pathological conditions. Superoxide radicals have very high rates of reactivity, which is why their excess damages many types of bio-molecules, including DNA. The mutations in Drosophilaflies' cells lowered their ability to resist that damage.
What was already known
According to the scientists, mutations in the genome of the experimental flies affected two genes: CG5017 and spineless. Both these genes are involved in regulation of many different functions, a large number of which still remain unknown. The team of researchers from MIPT, RAS and Cold Spring Harbor Laboratory has set a goal to reduce this white spot. The scientists started with the already known facts, i.e., that the CG5017 gene mutations impair insects' memory, and that mutations in spineless bring about a number of defects in morphogenesis. In Drosophila flies, spineless mutations affect leg and eye formation and disrupt the development of neuronal dendritic structure.
The CG5017 gene encodes the proteins which belong to the Nucleosome assembly protein (NAP) super-family. These proteins are the so-called nucleotropic chaperones, i.e., they are involved in the formation of functioning chromosomes' elementary units, namely, nucleosomes. It should also be noted that in most multicellular organisms the DNA molecules, located in the cell's nuclei, are not simply wound up double spirals, but very complex structures linked with protein molecules.
Spineless is a transcription factor, which means that this gene is involved in the process of transcribing genetic information from DNA to RNA, that is, RNA synthesis. This gene is important for cells also because it regulates the transcriptional activity of many other genes. The protein encoded by spineless regulates the synthesis of other proteins, which, in their turn, perform many important functions. Mutations appearing in spineless do not directly impact the structure of this or that protein. Yet, such mutations can either seriously limit, or even render impossible, the synthesis of many different protein molecules at once.
Spineless-controlled genes are responsible for the proper development of an animal. Even by the first half of the twentieth century,spineless gene mutations were ranked among the so-called homeotic mutations. Such mutations do not stop the process of the organism's formation, but direct it in the wrong way.
"Where we expected to see antennae, there emerged leg morphological structures with altered segmentation. It later turned out that the formation of eye structures and nervous cells had also been disrupted," said Boris Kuzin, the leading author of the article presenting the research. Boris Kuzin is a staff scientist of the DNBICT Brain Stem Cells Laboratory and the head of the Genetic Mechanisms of OrganogenesisResearch Group at the Institute of Developmental Biology (RAS).
What was discovered
The new research has shown the following: the spineless gene is also involved in long-term memory formation and detoxification. Spineless, and the proteins it controls, help the cell to get rid of exogenous toxins (i.e., originating outside the cell) and endogenous ones (in this case, those which were produced inside the cell following the irradiation). According to Boris Kuzin, the combination of the weak spineless and CG5017 gene mutations in a genome has a synergistic effect and results in the weakening of spineless (ss) products' functions. The organisms of fruit flies with these particular mutations cannot manage the simultaneous processes of detoxification and morphogenesis.
"It seems that the disruption of detoxification leads to aggravation of the defects, which appear in the process of limb formation in response to even low radiation doses. It looks like these mutant flies lack the functionally active spineless (ss) products necessary for the simultaneous control of development and detoxification," said Kuzin.
It should be emphasized that many genes have functional analogues, which work as partial substitutes, i.e., if a gene cannot function properly because of a mutation, other genes can, to some extent, compensate for this disruption. Besides, gene mutations can be strong (the affected gene is fully dysfunctional) and weak (the affected gene retains some of its functions).
What is means
The discovery, as the scientists state in their article, can shed some light on the problem of individual irradiation sensitivity. It is known that low doses of radiation sometimes result in serious inborn defects, and sometimes leave no traces. In part, it is connected with the a priori random nature of ionizing radiation, but there are also a number of genetically-based molecular-biological differences, many of which have not been yet defined.
"These results may have broader implications beyond the model organism. In particular, they may indicate an increased risk of pathological response to radiation in humans carrying hypomorphic mutations of these genes in their genome (note that both genes are highly evolutionarily conserved). Such individuals may be more vulnerable than the bulk of the population to even low levels of radiation, such as those delivered during routine medical procedures, prolonged air travel, or long term residency on the premises with high levels of radon," wrote the researchers. In the scientists' opinion, with the advent of broadly available genome sequencing it may become possible to single out the mutations linked with increased radiation sensitivity in individuals. The data can be used, for example, when choosing diagnostic procedures for such patients in order to ameliorate their increased vulnerability to radiation.
The fact that the CG5017 gene mutation effect is enhanced by a mutation in spineless (ss), and vice versa, confirms that different genes do interact with each other. The researchers suppose that the CG5017 gene product launches the process of synthesis of other proteins, which, in their turn, are necessary for the complete functioning of the spineless gene products.
A further detailed study of such interactions will help the scientists to get answers to many questions. For example, how multicellular organisms developed, how they acquired the ability to resist various toxins at different steps of evolution and how they managed to use this ability and long-term memory while adapting to a new ecological niche.
Researchers
The leading expert of the research project is Boris Kuzin, a staff scientist of the DNBICT Brain Stem Cells Laboratory and the head of the Genetic Mechanisms of Organogenesis Research Group at the Institute of Developmental Biology (RAS). Among Kuzin's scientific interests are gene transcription regulation processes, determination and differentiation of cells, and morphogenesis.
Grigori Enikolopov is the head of MIPT's DNBICT Brain Stem Cells Laboratory. Enikopolov is among the scientists awarded with the so-called "mega-grants", i.e., target financing as part of a Russian Federation government program, aimed at inviting leading specialists to universities.
Genes and what comes with them
DNA is able to store the information because its component blocks, the nucleotide pairs, are joined together in a certain order, the so-called nucleic acid sequence. The DNA matrix is the basis for the synthesis of another molecule, RNA. This molecule differs fromDNA in chemical content and structure: in most cases, RNA is a single, not a double, strand.
RNA molecules serve either as the matrix for protein synthesis (every three RNA nucleotides correspond to one amino acid in the protein), or perform their own function, e.g., some RNAs act as ferments, inducing biochemical reactions.
DNA, as was mentioned above, does not appear isolated in the cell, but always comes with a complex of proteins. These proteins are responsible for selectively "switching off" a certain number of genes in the cell. This, in turn, makes the process of RNA synthesis, ortranscription, impossible. There is also a group of proteins which, on the contrary, perform the function of launching RNA synthesis. Such proteins are called transcription factors.
Studying of the transcription control mechanisms (and today we know plenty of them) is the main task of molecular biology. There are many reasons for this.
First, it is gene transcription that defines the type of cells during the organism's development. Because of the differences in the processes of transcription, some cells become neurons, and others turn into liver, or, skin.
Second, understanding the differences in gene expression can shed light on the process of aging, the mechanisms of degradation of normal cells into cancer, and on many other disease progressions.
Third, during evolution, many important characteristics linked with the emergence of a new trait did not appear because of the arrivalof some absolutely new kind of protein. These characteristics were shaped by changes in the pattern of development of a certain organ. For example, our ancestors got rid of the tail obviously not because some new protein appeared, but because at a particular stage of embryonic development the group of genes responsible for tail vertebrae formation got "switched off".
Fourth, the switching "on" and "off" of the genes in a certain sequence supports many vital processes, including memory consolidation, digestion, muscle mass growth during long-term physical activity, and many others.
Many genes interact with each other, and this interaction can develop according to different scenarios. A protein encoded by one gene can engage with a protein encoded by another gene, and the resulting product, in its turn, will launch the RNA synthesis from a third, or a fourth, gene. Or, it might, on the contrary, suppress the synthesis of yet another protein.
These days the scientists are actively studying such interactions.
More information: Paper: www.plosone.org/article/info%3Adoi%2F10.1371%2Fjournal.pone.0094975















