Overlooked DNA shuffling drives deadly paediatric brain tumour
One of the deadliest forms of paediatric brain tumour, Group 3 medulloblastoma, is linked to a variety of large-scale DNA rearrangements which all have the same overall effect on specific genes located on different chromosomes. The finding, by scientists at the European Molecular Biology Laboratory (EMBL), the German Cancer Research Centre (DKFZ), both in Heidelberg, Germany, and Sanford-Burnham Medical Research Institute in San Diego, USA, is published online today in Nature.
To date, the only gene known to play an important role in Group 3 medulloblastoma was a gene called MYC, but that gene alone couldn't explain some of the unique characteristics of this particular type of medulloblastoma, which has a higher metastasis rate and overall poorer prognosis than other types of this childhood brain tumour. To tackle the question, Jan Korbel's group at EMBL and collaborators at DKFZ tried to identify new genes involved, taking advantage of the large number of medulloblastoma genome sequences now known.
"We were surprised to see that in addition to MYC there are two other major drivers of Group 3 medulloblastoma – two sister genes called GFI1B and GFI1," says Korbel. "Our findings could be relevant for research on other cancers, as we discovered that those genes had been activated in a way that cancer researchers don't usually look for in solid tumours."
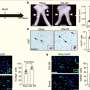
Rather than take the usual approach of looking for changes in individual genes, the team focused on large-scale rearrangements of the stretches of DNA that lie between genes. They found that the DNA of different patients showed evidence of different rearrangements: duplications, deletions, inversions, and even complex alterations involving many 'DNA-shuffling' events. This wide array of genetic changes had one effect in common: they placed GFI1B close to highly active enhancers – stretches of DNA that can dramatically increase gene activity. So large-scale DNA changes relocate GFI1B, activating this gene in cells where it would normally be switched off. And that, the researchers surmise, is what drives the tumour to form.
"Nobody has seen such a process in solid cancers before," says Paul Northcott from DKFZ, "although it shares similarities with a phenomenon implicated in leukaemias, which has been known since the 80s."
GFI1B wasn't affected in all cases studied, but in many patients where it wasn't, a related gene with a similar role, GFI1, was. GFI1B and GFI1 sit on different chromosomes, and interestingly, the DNA rearrangements affecting GFI1 put it next to enhancers sitting on yet other chromosomes. But the overall result was identical: the gene was activated, and appeared to drive tumour formation.
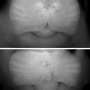
To confirm the role of GFI1B and GFI1 in causing medulloblastoma, the Heidelberg researchers turned to the expertise of Robert Wechsler-Reya's group at Sanford-Burnham. Wechsler-Reya's lab genetically modified neural stem cells to have either GFI1B or GFI1 turned on, together with MYC. When they inserted those modified cells into the brains of healthy mice, the rodents developed aggressive, metastasising brain tumours that closely resemble Group 3 medulloblastoma in humans.
These mice are the first to truly mimic the genetics of the human version of Group 3 medulloblastoma, and researchers can now use them to probe further. The mice could, for instance, be used to test potential treatments suggested by these findings. One interesting option to explore, the scientists say, is that highly active enhancers – like the ones they found were involved in this tumour – can be vulnerable to an existing class of drugs called bromodomain inhibitors. And, since neither GFI1B nor GFI1 is normally active in the brain, the study points to possible routes for diagnosing this brain tumour, too.
But the mice also raised another question the scientists are still untangling. For the rodents to develop medulloblastoma-like tumours, activating GFI1 or GFI1B was not enough; MYC also had to be switched on. In human patients, however, scientists have found a statistical link between MYC and GFI1, but not between MYC and GFI1B, so the team is now following up on this partial surprise.
"What we're learning from this study is that clearly one has to think outside the box when trying to understand cancer genomes," Korbel concludes.
More information: Northcott, P.A., Lee, C., Zichner, T. et al. Enhancer hijacking activates GFI1 family oncogenes in medulloblastoma. Published online in Nature on 22 June 2014. DOI: 10.1038/nature13379