New detection technique used in analysis of vaccines

(PhysOrg.com) -- An analysis of vaccines undertaken by researchers from 5 institutions has found that 7 of the vaccines' DNA content was pretty much as expected, but surprisingly, one also contained DNA of an apparently benign pig virus. This finding is reported in a paper written by lead author Eric Delwart of Blood Systems Research Institute and 6 co-authors, including three researchers from LLNL.
An analysis of vaccines undertaken by researchers from five institutions has found that seven of the vaccines’ DNA content was pretty much as expected, but surprisingly, one also contained DNA of an apparently benign pig virus.
This finding is reported in a paper written by lead author Eric Delwart of the San Francisco-based Blood Systems Research Institute and six co-authors, including three researchers from Lawrence Livermore National Laboratory (LLNL). The paper is published Wednesday in the online Journal of Virology.
Even before its publication, the research produced a major stir as Delwart contacted GlaxoSmithKline to alert the firm that its Rotarix vaccine, used to prevent diarrhea in babies, contained the pig virus, porcine circovirus-1 (PCV-1).
On March 22, Food & Drug Administration (FDA) officials urged pediatricians in the United States to temporarily suspend the use of Rotarix and to continue to use a similar rotavirus vaccine, RotaTeq from Merck, until further tests are completed. They added that Rotarix has been used in millions of children around the world, including one million in the United States, with no signs of safety problems, and that PCV1 is not known to cause any kind of illness in people or animals.
The European Union has opted for continued Rotarix distribution, while Glaxo removed PCV1 from their vaccine production line. The World Health Organization also recommended continued Rotarix distribution in regions with high levels of rotavirus infections in children.
“The intent in our research was to use the latest technologies to show that live vaccines only contained the expected viral genomes and no other,” Delwart said. “Indeed, seven of the eight vaccines were clean. We were quite surprised to find PCV-1 DNA in Rotarix.”
Delwart discovered PCV1 in Rotarix in January using a deep DNA sequencing technology and then contacted LLNL researchers to check his results.
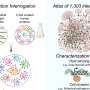
The three Livermore researchers - biologist Crystal Jaing, population biologist Shea Gardner and computer scientist Kevin McLoughlin - used a new LLNL detection technology, the Microbial Detection Array (MDA).
With 388,000 probes that fit on a one-inch wide, three-inch long glass slide, the MDA can detect or identify within a 24-hour period any of the approximately 60,000 viruses or 2,500 bacteria worldwide that have been sequenced.
“We have been collaborating with Eric for more than a year,” Jaing said. “He sees value in our microarray technology as a way to provide additional confirmation to his sequencing technology.”
The Livermore microarray confirmed the presence of PCV-1 DNA in the vaccine. The MDA has 40 designed probes to detect the PCV-1 strain of the pig virus; 39 of them showed positive for the virus, “almost a guarantee that the virus was present,” according to Jaing.
“One result of this research is that it demonstrates how modern technologies could change and drastically improve product safety,” said Tom Slezak, LLNL’s scientific leader for Bioinformatics.
As an example, Slezak noted that while current product safety rules require demonstrating that a list of known contaminants is not present, the use of modern advances in DNA sequencing and arrays would allow manufacturers to know every biological material that is present in quantities large enough to be of potential concern.
In their paper, the team wrote that while the polymerase chain reaction (PCR) technique is more sensitive than either metagenomic sequencing (where only a small sample is obtained of the total background DNA content of a complex mixture) or microarrays (such as the MDA) when screening for specific viruses, the sheer number of potential contaminating viruses limits the use of PCR to a small number of most commonly suspected viruses.
No one expected to see PCV-1 in a vaccine, so it was not on the list to be tested.
“The use of viral metagenomics and microarrays tests therefore seems warranted for the surveillance of products that are derived from cell cultures, plasma pools, or other biological sources and which may contain and transmit adventitious viruses,” the authors wrote.
Besides the Blood Systems Research Institute and LLNL, other institutions that collaborated on the Journal of Virology article include the Stanford Genome Technology Center, the David Grant U.S. Air Force Medical Center in Travis, Calif. and UC San Francisco’s Department of Laboratory Medicine.
In their research, Delwart’s team analyzed eight live attenuated viral vaccines - oral poliovirus, rubella, measles, yellow fever, varicella-zoster, measles/mumps/rubella and two rotavirus vaccines, one produced by Glaxo and the other by Merck.
Rotavirus can cause severe diarrhea and kills many children in developing nations. Before rotavirus vaccinations started in the United States, about 55,000 children per year were hospitalized for rotavirus infections and several dozen died each year. Merck introduced its vaccine in 2006 and Glaxo’s came out in 2008.
In their paper, the authors wrote that despite an extensive record of safety and efficacy, common misconceptions have continued about vaccine safety, leading to reduced childhood vaccinations and a resurgence of vaccine-preventable infections.
“Given that live attenuated viral vaccines are safe, effective and relatively inexpensive, their use against human and animal pathogens should be encouraged,” the authors said.
Application of new DNA technologies will reassure the public that viral contaminants can be readily detected and removed from vaccines, Delwart said.