Study discovers role of DNA methylation in multiple myeloma blood cancer
Sept. 30, 2010 — DNA methylation — a modification of DNA linked to gene regulation — is altered with increasing severity in a blood cancer called multiple myeloma, according to a study by Mayo Clinic and the Translational Genomics Research Institute (TGen).
And at specific points of DNA, "global hypomethylation," in which many genes lose the modification, may be associated with the step-by-step development of myeloma, according to a scientific paper published this month in the journal Cancer Research.
"This is the first study to show that hypomethylation occurs early in the development of multiple myeloma and increases through disease progression," said Dr. Bodour Salhia, a TGen cancer researcher and the paper's lead author.
DNA methylation suppresses the expression of viral genes and other harmful elements incorporated over time into an individual's genome. In cancer, hypermethylation at certain genomic locations can turn tumor suppressing genes off, while hypomethylation in some instances may lead to the over-expression of oncogenes, or those genes that give rise to cancer, and is linked to chromosomal instability.
However, there is still much to learn about the consequences of altered methylation.
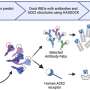
In this study, researchers examined the methylation status of more than 1,500 CpGs. This is shorthand for C-phosphate-G, or cytosine and guanine — two of the four chemicals that comprise DNA — separated by a phosphate group, which links the two nucleosides together.
Researchers used a high-throughput universal bead array technology to examine CpG methylation at different stages of multiple myeloma, evaluating DNA methylation events associated with the progression of tumors.
They performed DNA methylation profiling analysis for more than 800 genes, including tumor suppressors, oncogenes, and genes involved in cancer-related cellular processes. This process contrasts with previous studies that focused on the analysis of a single gene.
They found only a few genes that were hypermethylated, but importantly found many more hypomethylated genes, even in the earliest stages of multiple myeloma.
"Our data suggest that the overall degree of methylation may have some prognostic value, and further studies are needed to determine the functional and clinical significance of our findings," said Dr. John Carpten, Director of TGen's Integrated Cancer Genomics Division and the paper's senior author.
Dr. Salhia, added, "This study represents the most comprehensive examination to date of the role of methylation in multiple myeloma, and is expected to lead to an improved understanding of the biological mechanisms involved in the development of this type of cancer."
The study of DNA methylation falls under epigenetics — an emerging field in cancer research. Unlike the study of genetics, epigenetics refers to the study of gene activity that does not involve hardwiring alterations in the genetic code. These epigenetic events, which lay atop the genome, are an intricate and heritable mechanism of regulating the expression of genes.
"Understanding the full spectrum of epigenetic modifications will be key to improving the clinical management of the disease, and studies should continue to find new ways of treating multiple myeloma by targeting the multiple myeloma epigenome. This study also emphasizes that hypomethylating strategies may not be the next necessary steps in drug development." said Rafael Fonseca, M.D., Deputy Director of Mayo Clinic Cancer Center in Arizona.