'GPS system' for protein synthesis in nerve cells gives clues for understanding brain disorders
Scientists at the University of Pennsylvania explain how a class of RNA molecules is able to target the genetic building blocks that guide the functioning of a specific part of the nerve cell. Abnormalities at this site are in involved in epilepsy, neurodegenerative disease, and cognitive disorders. Their results are published this week in the journal Neuron.
A team of researchers, led by James Eberwine, PhD, the Elmer Bobst Professor of Pharmacology in the School of Medicine, and Junhyong Kim, PhD, the Edmund J. and Louise W. Kahn Professor of Biology in the School of Arts and Sciences, looked at how RNA gets targeted to nerve cell dendrites, which branch from the cell body of the neuron and detect the electrical and chemical signals transmitted by the axons of other neurons. These studies were enabled through the use of sensitive single cell analysis techniques developed in the Eberwine lab.
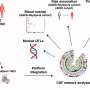
They discovered a class of RNAs (called CIRTs) that have small regions of retained strings of genetic building blocks (introns). These special RNAs have the ability to home to the dendrite to guide protein synthesis there. Specifically, they found that the targeting ability of some CIRTs originates from retrotransposons, which are thought to come from viruses.
The team concentrated on one retained intron, a localized regulatory sequence called the ID element. "Targeting elements in general are used by cells to make sure RNAs get to where they ultimately need to go, the dendrite in the case of a neuron. "It's like a GPS system for an RNA within the cell," says co-first author Peter T. Buckley, PhD, a postdoctoral fellow in the Eberwine lab. "But it gets removed in the cytoplasm before protein synthesis occurs. That's why the ID sequence isn't seen in the final protein."
The intron is a guide for local control of gene expression. The team used a reporter gene in the ID element to track its movement from the nucleus to the dendrite.
"There are species to species differences in the proportion and type of retrotransposons that make up introns," explains co-first author Miler T. Lee, PhD, a postdoctoral fellow in Junhyong Kim's lab. For example, rats have 100,000 of these ID elements versus the 1,000 to 2,000 found in mice. "Our studies suggest that rats use this ID element to target mRNA to the dendrite while mice may use other localization mechanisms." These data suggest that researchers must be careful in selecting animal models for the study of neurological and psychiatry illnesses.
One of the ID elements the researchers identified and analyzed is in an intron present in the FMRI gene. Fragile X syndrome -- one of the most common causes of inherited mental retardation -- is caused by mutations in this gene. The gene encodes the FMRP protein, which controls the availability of select proteins involved in neuron-to-neuron communication. The ID element in the FMR1 mRNA, in part, targets the RNA to where the FMRP protein is synthesized, which in turn controls how and where other proteins are made.
Because some retrotransposons are derived from viruses these data provide a mechanism by which normal cell functioning could be altered by viral infection. Upon infection of cells by viruses cellular proteins are often hijacked to permit the virus to function and divide. Since some RNAs that are involved in learning and memory contain ID sequences very similar to viruses, perhaps the proteins that move the RNAs to the dendrite are hijacked by an invading virus, surmise the researchers. The result being that the normal cell RNA does not move to its proper cellular position. If this happens during a critical period of development proper neuronal connectivity may be compromised and could result in long-term dysfunction of the central nervous system. This research area is currently under active investigation in the Eberwine and Kim labs.
These intron sequences have previously been thought to be junk RNA with no function, yet as shown in this study -- in some cases -- introns are functional outside of the nucleus. Extensive bioinformatic analysis performed by the team also suggests other functions including the possibility that outside of the nucleus retained introns may produce small RNAs that could regulate other RNAs.
The teams are now looking into perturbing this intricate system to see if they can change RNA targeting, as well as translation and protein function.