Precision gene targeting in stem cells corrects disease-causing mutations
Using two distinct methods, Whitehead Institute researchers have successfully and consistently manipulated targeted genes in both human embryonic stem (ES) cells and induced pluripotent stem (iPS) cells (adult cells that have been reprogrammed to an embryonic stem cell-like state).
In one case, scientists employed proteins known as zinc finger nucleases (ZFNs) to change a single base pair in the genome, allowing them either to insert or remove mutations known to cause early-onset Parkinson's disease (PD). The second method relies on proteins called transcription activator like effector nucleases (TALENs) capable of altering specific genes with similar efficiency and precision as ZFNs. Both sets of experiments were conducted in close collaboration with scientists at Sangamo BioSciences.
Targeted genetic manipulation addresses a problem that has been plaguing human stem cell research – the ability to cleanly and site-specifically modify the genomes of human ES and iPS cells. Realizing the therapeutic promise of these cells depends on such changes to fix disease-causing mutations before the cells could be transplanted into patients or to create cell lines that researchers can use to study genetic diseases.
Such disease studies—the much-heralded "disease in a dish" approach—and the search for potentially disease-modifying drugs require the use of cells and controls that are genetically identical, except for a specific alteration whose impact can then be observed.
"This is very relevant for diseases like Parkinson's, which likely will display only subtle phenotypes in the Petri dish. It is very important that the cells be genetically identical and have the same history, then make or remove only that mutation," says Whitehead Founding Member Rudolf Jaenisch. "If you use control cells from one person and a diseased cell from another person, it's like comparing apples and oranges."
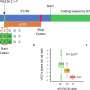
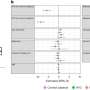
As reported in a paper published July 22 in Cell, first author Frank Soldner used ZFNs created by Sangamo BioSciences to generate, from both normal and PD patients' cells, sets of mutated and control cell lines. By either removing or adding a mutation to the alpha-synuclein gene associated with PD, Soldner created lines of cells whose genomes differ only by a single base pair. Subsequent differences seen in comparative studies of the cells can therefore be attributed to the mutation in question.
"ZFNs can transfer a mutation without any other alterations to the genome, such as leaving in unwanted pieces of DNA that could be harmful," says Soldner, a postdoctoral researcher in Jaenisch's lab. "This precision is ideal for drug research for PD and other diseases, but it is also one more step toward using ES or iPS cells therapeutically."
In its continual quest to refine human stem cell technology, the Jaenisch lab has also been investigating other gene targeting approaches. One option is to use TALENs, which use a type of DNA-binding domain originally found in some plant pathogens. TALENs can be designed and created in academic labs.
To compare TALENs' ability to alter genes to that of ZFNs', two postdoctoral researchers in Jaenisch's lab, Dirk Hockemeyer and Haoyi Wang, repeated an earlier ZFN experiment, this time using TALENs created by scientists at Sangamo BioSciences. In research reported earlier this month in Nature Biotechnology, Hockemeyer and Wang show that these TALENs can also modify genes as efficiently and precisely as ZFNs in ES and iPS cells.
"These are amazing proteins," says Wang. "In theory, everything ZFNs do, they should be able to do as well."
"This opens up a lot of possibilities of what we will be able do because the generation of TALENs is extremely versatile," adds Hockemeyer. "It appears they, along with ZFNs, will help us overcome the challenges of developing human ES and iPS cell technology."
More information:
"Generation of isogenic pluripotent stem cells differing exclusively at two early onset Parkinson point mutations," Cell, July 22, 2011.
"Genetic engineering of human pluripotent cells using TALE nucleases," Nature Biotechnology, July 7, 2011.