New software opens the door to wider use of 3-D imaging in the study of disease
Researchers have developed a novel, easy-to-use system for three-dimensional (3D) reconstruction and examination of tissues at microscopic resolution, with the potential to significantly enhance the study of normal and disease processes, particularly those involving structural changes. The new approach, using conventional histopathological methods, is described in the May issue of The American Journal of Pathology.
"The use of 3D imaging technology to study structure, function, and disease manifestations has been limited because of low resolution, and the time and difficulty associated with acquiring large numbers of images with a microscope," says lead investigator Dr. Darren Treanor, University of Leeds and the Leeds Teaching Hospitals NHS Trust, United Kingdom. "Our system can integrate tissue micro-architecture and cellular morphology on large tissue samples. It can be used by technical or medical staff in a histopathology laboratory without input from computing specialists."
Developed by Dr. Derek Magee at the University of Leeds, the system utilizes automated virtual slide scanners to generate high-resolution digital images and produce 3D tissue reconstructions at a cellular resolution level and can be used on any stained tissue section. It is based on a general image based-registration algorithm and operates using an integrated system that requires minimal manual intervention once the slides are sectioned, stained, and mounted. The virtual slide scanners digitize the tissue automatically, the software communicates with the software serving the image, which aligns the images, and produces visualization in one integrated package. The user can manually select a region, zoom in and re-register the area to get a higher resolution image of microscopic features.
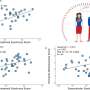
The authors have applied the system to over 300 separate 3D volumes from eight different tissue types, using a total of 5,500 virtual slides. They describe cases that illustrate the possible applications of the system. For example, a 3D volume rendering of a mouse embryo demonstrates that the method could be useful for providing anatomical and expression data and for creating a "virtual archive" of 3D transgenic models. A 3D volume rendering of sections from a human liver containing a deposit of metastatic colorectal carcinoma adjacent to a blood vessel could provide insight into tumor vasculature and its response to anti-angiogenic agents. A 3D visualization of cirrhotic human liver infected with hepatitis C demonstrates the software's potential to provide information on disease development and aid diagnosis.
"Many fields, including tumor biology, embryology, and cardiovascular disease could benefit from correlation of structure and function in three dimensions, but getting high quality 3D reconstructions has always been difficult" says Dr. Treanor. "We have demonstrated that our software is accurate and robust enough to use without significant computer science input. This system provides the opportunity for increasing use of 3D histopathology as a routine research tool."