Advanced cancers destined to recur after treatment with single drugs that 'target' tumor cells: study
Targeted cancer cell therapies using man-made proteins dramatically shrink many tumors in the first few months of treatment, but new research from Johns Hopkins scientists finds why the cells all too often become resistant, the treatment stops working, and the disease returns.
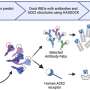
In a study of 28 advanced colon cancer patients treated with the monoclonal antibody panitumumab, the Johns Hopkins Kimmel Cancer Center team reports that drug-resistance tumor cell mutations appear in the blood of patients five to seven months later, and that low levels of these mutations exist in nearly all tumors before the therapy begins, making the cancers predestined to recur.
"These resistance mutations develop by chance as cancer cells divide so that tumors always contain thousands of resistance cells," says Luis Diaz, M.D., associate professor of oncology and director of the Swim Across America laboratory at Johns Hopkins, who says the findings likely apply to any targeted cancer therapy.
"The best chance for a cure is when a tumor is very small, but when the cancer is advanced, our research quantifies the probability that we can achieve cures with single-agent targeted therapies," says Bert Vogelstein, M.D., professor and co-director of the Ludwig Center at Johns Hopkins and, Howard Hughes Medical Institute investigator. "Long-term remissions of advanced cancers will be nearly impossible with single targeted agents," he adds.
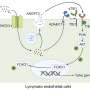
The Johns Hopkins scientists analyzed blood samples taken from 28 patients with advanced colorectal cancers. These patients were enrolled in a clinical trial of panitumumab, one of a new and growing class of monoclonal antibodies, or synthetic proteins that homes in on cancer cells' vital growth pathways. In the case of panitumumab, the agent targets a growth-factor receptor called EGFR. Patients most likely to respond to the drug also have normal copies of the KRAS gene in their tumors.
Twenty-four of the 28 patients in the study had normal KRAS gene copies in their tumors, and four had mutations in KRAS, serving as a control group. Blood samples were taken before beginning the therapy and at four-week intervals during the therapy, for a total of 169 combined blood draws.
Virtually all cancers shed DNA material into the blood, according to the researchers, and provide an easy route to collecting molecular evidence from lesions typically inaccessible for surgical biopsy. "The amount of tumor DNA found in the blood is akin to tests used to determine HIV viral load," says Diaz.
In their analysis, reported online June 13 in the journal Nature, the scientists found that nine of the 24 patients with normal KRAS genes (38 percent) exhibited KRAS mutations detectable in the blood within five to seven months of beginning therapy. KRAS mutations were detected in three patients before imaging scans showed metastatic tumor growth. Then, working with Martin Nowak, Ph.D., and his team from Harvard University, the investigators used mathematical models to calculate when KRAS mutations likely originated. Nowak and colleagues determined that KRAS mutations were present prior to the initiation of treatment with panitumumab.
"The probability that the mutations were absent at the beginning of treatment is exceedingly low," says Vogelstein, leading the team to conclude that the development of drug-resistance is a fait accompli. The time it takes for cancers to recur is determined simply by how long it takes cancer cells with mutant genes to multiply, he adds.
The research team says that combination therapies are the best chance for longer remissions. "The good news is that there is a limited number of pathways that go awry in cancer, so it should be possible to develop a small number of agents that can be used in a large number of patients," says Vogelstein. "However, I hope this research will help stimulate the testing of new drugs as combination therapies much earlier in the drug approval process than the current norm."
More information: DOI: 10.1038/nature11156 , DOI: 10.1038/nature11219