From genome to immune response

Being able to predict which peptides are more efficient at generating immune responses would be ideal when designing vaccination approaches. European scientists made this possible by developing state-of-the-art bioinformatics tools.
With the advent of genome sequencing technologies, it has become easier to sequence the entire genome of pathogens in a short period of time. Such information could be used to study the biology of almost any pathogen and its potential effect – through the translated proteins – to the immune system.
Peptides are among the key processing units of our immune system. T cells have been developed to be able to recognise and bind peptides presented on MHC molecules of antigen-presenting cells or host cells. Peptide generation and presentation is a rather complicated process involving many steps. In essence, each individual presents to their T cells a unique and highly diverse peptide imprint of the ongoing protein metabolism.
The rationale behind the EU-funded ‘Translating genome and proteome information into immune recognition’ (Genomes_TO_Vaccines) project was to understand, describe and predict how the immune system handles proteins.
Efficient biochemical assays were initially developed to be able to evaluate the three major steps in the generation of T cell epitopes. These included the proteolytic fragmentation of protein antigens in the proteasome, the peptide translocation event (TAP), and the peptide selection and presentation on MHC molecules.
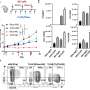
Scientists used bioinformatics to predict the outcome of these processes in silico and potential human T cell responses. Influenza A viral isolates were used to test the epitope scanning process, and predict peptides both in biochemical MHC binding assays as well as, more importantly, in healthy influenza convalescents.
Combined with data on the SARS virus, the Genomes_TO_Vaccines study identified 13 new influenza epitopes, which covered virtually the entire population. These epitopes were selected for being highly conserved and, notably, they all encompassed the current bird flu isolates.
Genomes_TO_Vaccines demonstrated the feasibility of translating genomic information into immune recognition. The developed technology has the potential to be utilised for designing immunotherapy and vaccination strategies, thus improving the quality of life for European citizens.