November 23, 2012 report
Proteins expressed by human cytomegalovirus mapped
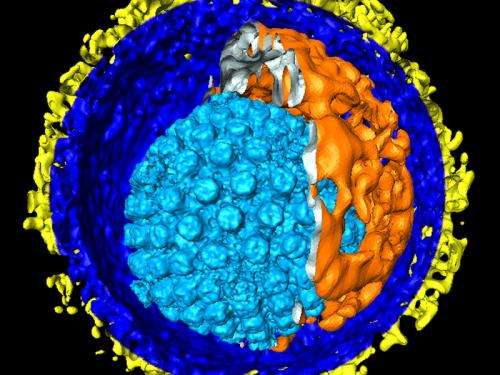
(Medical Xpress)—A new study in the US and Germany has added to our understanding of the human cytomegalovirus (HCMV) and how it manipulates the cells it infects.
HCMV is a very common herpes virus infecting 50-100% of adults, usually with no ill-effects, and it can remain latent for many years. HCMV infections are more common in people in developing countries than in advanced countries. The virus can cause birth defects in newborns and can cause serious and even fatal disease in people with compromised immune systems. The virus has also been implicated as possibly having a role in cancer development.
The new research by Noam Stern-Ginossar of the Jonathan Weissman laboratory at the University of California San Francisco and a team of US and German colleagues looked at the virus's proteome, which describes all the expressed proteins. The researchers thought the maps of potential proteins coded for by the genome, which were largely on computer models, were incomplete.
The HCMV genome was first sequenced over two decades ago and since then 16 more strains of the virus have been sequenced. It is a particularly large viral genome, with approximately 240,000 base pairs.
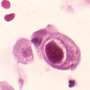
The team began by mapping the positions of the ribosomes during an HCMV infection of human fibroblast cells. The ribosomes are the organelles within cells in which proteins are manufactured. They then used mass spectrometry to determine if the predicted proteins were present. The research identified codings in the genome for hundreds of proteins that had not been previously identified.
The DNA segments encoding for the proteins are called "open reading frames," and previous research had predicted there would be 200 frames, based on an analysis of the genome sequences. In the new study the team found that some of these frames encoded extremely short protein sequences comprising 100 amino acids or fewer. They also found that some of the open reading frames were inside other frames. The complexity they found in the proteome was surprising and unexpected.
An understanding of what proteins the virus genome can express could help in determining how the virus affects the host cell, and how its manipulation of the cell could be disrupted.
In their paper, published in Science, the research team said their method of combining ribosome profiling and mass spectrometry could be used to examine other complex virus proteomes.
More information: Decoding Human Cytomegalovirus, Science, 23 November 2012: Vol. 338 no. 6110 pp. 1088-1093. DOI: 10.1126/science.1227919
ABSTRACT
The human cytomegalovirus (HCMV) genome was sequenced 20 years ago. However, like those of other complex viruses, our understanding of its protein coding potential is far from complete. We used ribosome profiling and transcript analysis to experimentally define the HCMV translation products and follow their temporal expression. We identified hundreds of previously unidentified open reading frames and confirmed a fraction by means of mass spectrometry. We found that regulated use of alternative transcript start sites plays a broad role in enabling tight temporal control of HCMV protein expression and allowing multiple distinct polypeptides to be generated from a single genomic locus. Our results reveal an unanticipated complexity to the HCMV coding capacity and illustrate the role of regulated changes in transcript start sites in generating this complexity.
© 2012 Medical Xpress