New discovery in quest for better drugs

Scientists have combined cutting edge computer modelling with pharmacology and medicinal chemistry to reveal new insights into how the body interacts with novel drug treatments, in research that could lead to the creation of drugs that are more targeted and with fewer side effects.
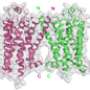
In a paper published today in Nature, researchers from the Monash Institute of Pharmaceutical Sciences (MIPS) were part of an international team who investigated alternative drug recognition sites on G protein-coupled receptors (GPCRs) - the largest and most important family of receptor proteins in the human body.
GPCRs play a role in virtually every biological process and most diseases, including neuropsychiatric disorders, cardiovascular disease, obesity and diabetes, inflammation and cancer. Almost half of all current medications available use GPCRs to achieve their therapeutic effect.
The new research into how GPCRs work at the molecular level has unlocked vital insights into how drugs interact with this therapeutically relevant receptor family.
Professor Arthur Christopoulos from MIPS said it was hoped the research would lead to the creation of drugs that are more targeted, and with fewer side effects.
"This study has cracked the secret of how a new class of drug molecule, which we have been studying for some time now, actually binds to a GPCR and changes the protein's structure to achieve its unique molecular effect," Professor Christopoulos said.
"This research can explain the behaviour of such drugs at the molecular level and facilitate structure-based design for new and more potent drugs."
By starting with a known crystal structure of a GPCR as a template, the team used computer simulations to map how different drugs and the receptor can "find" each other, and how they change their shape and orientation as they interact. Importantly, the predictions made by the computer simulations were validated by new biological experiments and by the rational design of a more potent molecule that targets the GPCR.
More information: www.nature.com/nature/journal/ … ull/nature12595.html