Gene-silencing strategy opens new path to understanding Down Syndrome
The first evidence that the underlying genetic defect responsible for trisomy 21, also known as Down syndrome, can be suppressed in laboratory cultures of patient-derived stem cells was presented today (Oct. 22) at the American Society of Human Genetics 2013 annual meeting in Boston.
People with Down syndrome are born with an extra chromosome 21, which results in a variety of physical and cognitive ill effects. In laboratory cultures of cells from patients with Down syndrome, an advanced genome editing tool was successfully used to silence the genes on the extra chromosome, thereby neutralizing it, said Jeanne Lawrence, Ph.D., Professor of Cell & Developmental Biology at the University Massachusetts Medical School, Worcester, MA.
Dr. Lawrence and her team compared trisomic stem cells derived from patients with Down syndrome in which the extra chromosome 21 was silenced to identical cells from patients that were untreated. The researchers identified defects in the proliferation, or rapid growth, of the untreated cells and the differentiation, or specialization, of untreated nervous system cells. These defects were reversed in trisomic stem cells in which the extra chromosome 21 was muted.
"Silencing of trisomy 21 by manipulation of a single gene in living cells in laboratory cells surmounts the first major obstacle to development of potential 'chromosome therapy,'" said Dr. Lawrence, whose presentation today provided an update to the results that she and her colleagues reported earlier this year in the journal Nature (Jiang et al. 2013)
In her ASHG presentation, Dr. Lawrence described the use of the novel editing tool to examine changes in gene expression that result from the silencing of the extra chromosome. The changes in gene expression were not limited to chromosome 21 but were genome-wide.
"In fact, the results indicate that the most prominent changes are in genes not encoded on chromosome 21," said Dr. Lawrence, who also provided more perspective about the various avenues of research that the results have created and that are now being and will be pursued in her lab.
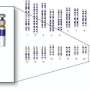
The approach used by Dr. Lawrence and her team was inspired by the natural process that silences one copy of the female mammals' two sex-determining X chromosomes during embryonic development. In males, the sex-determining chromosomes are X and Y, and gene silencing helps maintain similar expression patterns of X chromosome genes in females and males.
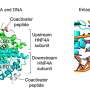
To understand this biological process, Dr. Lawrence and her collaborators several years ago began studying the X-inactivation gene (XIST), which encodes a large non-coding RNA molecule. In laboratory cultures of cells, this molecule was shown to cover the surface of one of the X chromosomes of female mammals. XIST's actions permanently blocked the expression, or activity level, of the genes on the affected X chromosome.
Dr. Lawrence and her team mimicked this natural process by inserting the XIST gene into the gene-rich core of the extra chromosome 21 in lab cultures of pluripotent stem cells from patients with Down syndrome. Before taking this step, they first demonstrated that a large transgene could be successfully inserted at a specific site by using zinc-finger nuclease technology.
In the laboratory cells, they found that the RNA from the inserted XIST gene induced a host of epigenetic modifications that transcriptionally silenced the genes of the extra chromosome 21.
"Remarkably, the RNA localized across and comprehensively silenced one of the three chromosome 21s, as shown by eight different methods, including molecular, cytological and genomic," said Dr. Lawrence.
The researchers found that APP, which encodes beta-amyloid precursor protein, was among the silenced genes on the XIST-coated chromosome 21. Mutations in APP cause the accumulation of beta-amyloid that leads to early-onset familial Alzheimer's disease, Dr. Lawrence said. APP over-expression is linked to the Alzheimer's disease that occurs in many patients with Down syndrome.
"The results show the clear promise of this new strategy as a novel approach to identify the poorly understood cellular pathways deregulated in Down syndrome and creates the opportunity to derive and study various patient-compatible cell types potentially relevant to Down syndrome therapeutics," she noted.
"This general strategy could be extended to study other chromosomal disorders such as trisomy 13 and 18, which are usually fatal in the first one to two years of life," Dr. Lawrence added.
More information: Title of the abstract: "Translating dosage compensation to Trisomy 21: a novel approach to Down syndrome."