Making the most of what you have

Understanding how viral proteins are produced can provide important clues on how we might interfere with the process. The group of Till Rümenapf at the University of Veterinary Medicine Vienna has discovered that a key protease of a particular virus breaks itself down into two different functional molecules. The findings are reported in the Journal of Virology and may have important implications for the development of defense strategies against diseases caused by flaviviruses.
We generally do not devote much time to worrying about the problems faced by viruses, although understanding them may provide clues on how to combat diseases. Viruses rely on the machinery of the host cell but must supply specific functions via their own proteins and RNA molecules. All of these need to be coded in as little genetic material as possible, so it is often the case that a single viral protein is able, indeed required, to undertake a number of different tasks. And the requirement to minimize the amount of DNA or RNA often leads viruses to lump all their proteins in a single gene, encoding what is known as a polyprotein. Polyproteins are a typical feature of RNA viruses, such as flaviviruses.
Flaviviruses in animals and man
Flaviviruses are responsible for a wide range of diseases in humans (such as tick borne encephalitis, dengue fever and hepatitis C) and animals (such as bovine viral diarrhoea and classical swine fever). Understanding how they function represents a key first step in the development of effective countermeasures or cures, so work on flaviviruses forms an import focus of a large number of research institutions worldwide.
The University of Veterinary Medicine Vienna has been contributing to the field since the appointment in 2012 of Till Rümenapf.
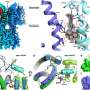
Rümenapf works on classical swine fever virus, CSFV, an RNA virus that encodes a single protein that must be broken down into twelve building blocks before the virus can exert any of its unwelcome effects on pigs. Key among the "mature proteins" are those responsible for chopping off the various pieces to produce functional proteins. The so-called "nonstructural protein 3" (NS3) appears to have a crucial role: not only is its protease activity important in the production of a number of other viral proteins, it also harbours further functions that are essential for replicating the viral RNA and for the formation of the virus particle.
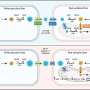
Benjamin Lamp and colleagues in Rümenapf's group have investigated NS3 in more detail. Surprisingly, they found that NS3 also attacks itself, generating two smaller proteins. The fractions are not merely breakdown products – proteases can naturally digest themselves – but proteins with distinct activities, a protease and a helicase (which is involved in RNA replication). The protease can actually cleave itself a further time, cutting off a small portion and leaving the remainder inactive.
Self-cleavage of NS3 is essential
The NS3 protease cuts proteins, even itself, only at precisely defined positions. Lamp altered the two newly detected cleavage sites and examined the effect on the virus. Changes at either site were found to cause dramatic drops in the efficiency of viral replication, strongly suggesting that both cleavages are important for the viral life-cycle, although what the resulting sub-proteins do is still unclear. As Rümenapf says, "People have been actively studying the NS3 protein for about twenty years so we were amazed to find new cleavage sites. The knowledge that NS3 is auto-cleaved changes the way we think about how the virus works and could be very helpful in understanding the complex regulation of viral replication that – in some flaviviruses such as hepatitis C virus and CSFV – leads to persistent infections."
More information: The paper "Autocatalytic Cleavage within Classical Swine Fever Virus NS3 Leads to a Functional Separation of Protease and Helicase" by Benjamin Lamp, Christiane Riedel, Eveline Wentz, Maria-Alejandra Tortorici and Till Rümenapf has just been published in the Journal of Virology (DOI: 10.1128/JVI.00754-13). jvi.asm.org/content/87/21/11872#ref-list-1