Sieving through 'junk' DNA reveals cancer-causing genetic mutations
Researchers can now identify DNA regions within non-coding DNA, the major part of the genome that is not translated into a protein, where mutations can cause diseases such as cancer.
Their approach reveals many potential genetic variants within non-coding DNA that drive the development of a variety of different cancers. This approach has great potential to find other disease-causing variants.
Unlike the coding region of the genome where our 23,000 protein-coding genes lie, the non-coding region - which makes up 98% of our genome – is poorly understood. Recent studies have emphasised the biological value of the non-coding regions, previously considered 'junk' DNA, in the regulation of proteins. This new information provides a starting point for researchers to sieve through the non-coding regions and identify the most functionally important regions.
"Our technique allows scientists to focus in on the most functionally important parts of the non-coding regions of the genome," says Professor Mark Gerstein, senior author from the University of Yale. "This is not just beneficial for cancer research, but can be extended to other genetic diseases too."
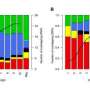
The team used the full set of genetic variants from the first phase of the 1000 Genomes Project, together with information about the non-coding regions generated by the ENCODE Project, and identified regions that did not accumulate much variation. Protein-coding genes play a crucial role in human survival and fitness, and are under strong 'purifying' selection, which removes variation. The team found that some non-coding DNA regions showed almost the same low levels of variation as protein-coding genes, and called these 'ultrasensitive' regions.
Within the ultrasensitive regions, they looked at specific single DNA letters that, when altered, caused the greatest disturbance to the genetic region. If this non-coding, ultrasensitive region is central to a network of many related genes, variation can cause a greater knock-on effect, resulting in disease.
They integrated all this information to develop a computer workflow known as FunSeq. This system prioritises genetic variants in the non-coding regions based on their predicted impact on human disease. "Our method is a practical and successful way to screen for purifying selection in non-coding regions of the genome using freely available data such as those from the ENCODE and 1000 Genomes Projects," says Dr Yali Xue, author from the Wellcome Trust Sanger Institute. "It really shows the value of these large-scale open access data-sets."
The team applied FunSeq to 90 cancer genomes including breast cancer, prostate cancer and brain tumours, and found nearly 100 potential non-coding cancer driving variants. In the breast cancer genomes, for example, they found a single DNA letter change that seems to have great impact on the development of breast cancer. This single letter change occurs in an ultrasensitive region that is central to a network of many related genes.
"Although we see that the first effective use of our tool is for cancer genomes, this method can be applied to find any potential disease-causing variant in the non-coding regions of the genome," says Dr Chris Tyler-Smith, lead author from the Wellcome Trust Sanger Institute. "We are excited about the vast potential of this method to find further disease-causing, and also beneficial variants, in these crucial but unexplored areas of our genome."
More information: Ekta Khurana, Yao Fu, Vincenza Colonna, Xinmeng Jasmine Mu et al (2013). "Integrative annotation of variants from 1,092 humans: application to cancer genomics" Advanced online publication in Science, 03 October, 2013.