Can thermodynamics help us better understand human cancers?

(Medical Xpress)—When the "war on cancer" was declared with the signing of the National Cancer Act in 1971, identifying potential physical traits, or biomarkers, that would allow doctors to detect the disease early on was a significant goal. To this day, progress in the battle against cancer depends on understanding the underlying causes and molecular mechanisms of the disease.
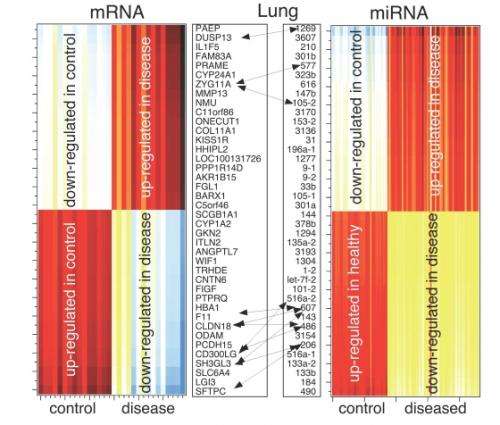
In a new study, UCLA researchers analyzed the gene-expression profiles of more than 2,000 patients and were able to identify cancer-specific gene signatures for breast, lung, prostate and ovarian cancers. The study applied an innovative approach to gene-array analysis known as "surprisal analysis," which uses the principles of thermodynamics—the study of the relationship between different forms of energy—to understand cellular processes in cancer.
The research appears in the early online edition of Proceedings of the National Academy of Sciences and will be published in an upcoming print edition.
Surprisal analysis allows researchers to observe how cellular energy is expended in cancer cells and how this process affects the way in which these cells choose to express certain genes. In particular, scientists can look at how cancer cells decide to use energy when expressing critical genes that allow them to persist and grow.
By identifying such cancer-specific gene signatures, scientists are able to distinguish, with high fidelity, the biopsy samples of cancer patients from control samples and potentially to identify novel cancer biomarkers for early detection of the disease and the development of new therapies.
Research co-author Raphael Levine, a UCLA distinguished professor of chemistry and biochemistry and of medical and molecular pharmacology, and his fellow researchers hope the cancer-specific signatures they identify using surprisal analysis will provide "thermodynamic targets" against cancer.
"We believe that this paper introduces a new hallmark of cancer—a thermodynamic signature—where the free energy redistributions among cellular biomolecules in the cancer state, not seen in the non-cancer state, sustain the disease," said Levine, a faculty member in the UCLA College of Letters and Science. "A further, future power of surprisal analysis is in its ability to detect 'patient potentials,' meaning patient-specific differences can be detected in the analysis, reintroducing the possibility of personalized medicine to the cancer arena."
The same year the war on cancer was declared, Levine first formulated surprisal analysis with his colleagues, the late Richard Bernstein of UCLA and Avinoam Ben-Shaul, recognizing a need to better understand and characterize how systems utilize energy. Since then, surprisal analysis has become a critical tool for the analysis of chemical, nuclear and physical dynamics. For this research, Levine was awarded the prestigious Wolf Prize in chemistry in 1988.
In the last few years, Levine and his colleges have attempted to extend surprisal analysis to biological systems. Because the theoretical approach enables the monitoring of small systems that are not in thermodynamic equilibrium, living cancer cells provide a very suitable opportunity for study.
Progress toward personalized medicine is expected to transform the cancer therapeutics field. But identifying an approach that can give researchers a feasible, quantitative method to identify cancer-specific gene signatures and characterize individual patient cancers has remained "an ultimate challenge," said Levine, a member of the U.S. National Academy of Sciences.
The new research suggests there may be an effective approach to identify cancer-specific signatures. While encouraging, Levine and his co-authors emphasize that the results are preliminary and that any practical diagnostics resulting from this research would require considerable additional and extensive scientific validation.
Cancer deaths have declined only slightly in the past decade, mostly due to preventive health efforts such as smoking cessation and routine examinations. Identifying cancer-specific markers remains a challenge. However, modern genetic sequencing technology can measure the expression levels of many genes. The recent development of large-scale genomic approaches and sequencing initiatives have produced several candidate biomarkers for cancer detection, but very few have been robust enough to work well in practice. The inability to extend these biomarkers from the bench to the clinic arises from a limited ability to separate the wheat from the chaff while sifting through the often-insurmountable data retrieved from genomic technologies, Levine said.
Co-authors of the study are Francoise Remacle, a professor of chemistry at the University of Liege in Belgium and director of research at the Belgian National Scientific Research Foundation (FNRS), and lead author Sohila Zadran, a junior faculty fellow at the David Geffen School of Medicine at UCLA.
More information: www.pnas.org/content/early/201 … 991110.full.pdf+html