January 23, 2014 report
Team develops method of identifying impact of gut microbes
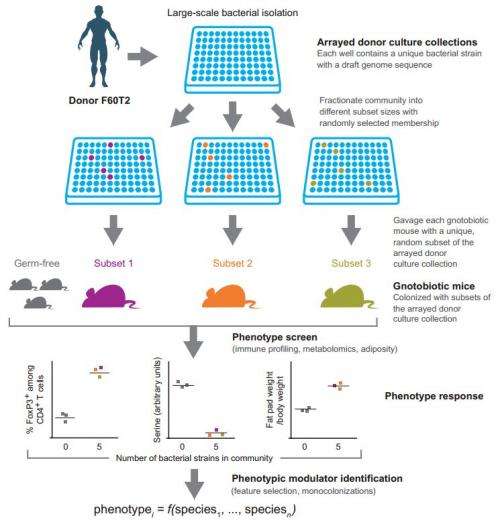
(Medical Xpress)—A team of researchers at Washington School of Medicine in St. Lois has developed a method for identifying the impact that individual strains of microbes in the human gut have on the person housing them. In their paper published in the journal Science Translational Medicine, the team describes how they isolated different microbes taken from human volunteers and placed them in the guts of sterilized mice, and then tested them to see what impact it had.
Over the past few decades, scientists have come to realize that microbes in our guts play a far more important role in our lives than anyone might have ever thought possible. Scientists now know that in addition to helping us ward off diseases and digest our food, microbes are also in part responsible for how much fat our bodies hold onto and for causing ailments such as colitis, Irritable Bowel Syndrome or Crohn's Disease.
As more has become known about the impact of microbes in our gut, it's become clear that a means for figuring out which do what would be very helpful for treating ailments or even for helping lose weight. The problem with doing so is there are trillions of such microbes in our guts, with perhaps 10,000 different species. Undaunted, the researchers with this new effort have developed a method for doing so anyway.
In order to understand what a microbe does, the researchers reasoned, it must be isolated and allowed to do whatever it does under close scrutiny. To achieve that goal, they collected fecal specimens from several human volunteers, took them to a lab and separated out 94 different microbe varieties. Each was then introduced individually into the gut of a mouse that had had its gut cleared of all microbes. Thus, the team was able to monitor the impact of each single microbe on the mouse being tested.
In so doing, the team discovered that introducing one of eight types of microbes into their guts caused the mice to gain weight. A mouse with no gut bacteria is generally very thin, as it gets no help with digesting food, thus the bacteria introduced have now been identified as being among those that impact fat retention—a development that could at some point result in a probiotic pill for humans that have trouble digesting certain foods.
The method is slow of course, but so is a lot of scientific research. The point is that the process appears to be one that will allow researchers over time to identify which gut microbes do what. That won't reveal all of course, as scientists know that sometimes some impacts on the body are the result of intermingling of microbes, but it appears to be a great start in mapping human gut microbes.
More information: J. J. Faith, P. P. Ahern, V. K. Ridaura, J. Cheng, J. I. Gordon, Identifying Gut Microbe–Host Phenotype Relationships Using Combinatorial Communities in Gnotobiotic Mice. Sci. Transl. Med. 6, 220ra11 (2014). stm.sciencemag.org/content/6/220/220ra11
Abstract
Identifying a scalable, unbiased method for discovering which members of the human gut microbiota influence specific physiologic, metabolic, and immunologic phenotypes remains a challenge. We describe a method in which a clonally arrayed collection of cultured, sequenced bacteria was generated from one of several human fecal microbiota samples found to transmit a particular phenotype to recipient germ-free mice. Ninety-four bacterial consortia of diverse size, randomly drawn from the culture collection, were introduced into germ-free animals. We identified an unanticipated range of bacterial strains that promoted accumulation of colonic regulatory T cells (Tregs) and expansion of Nrp1lo/− peripheral Tregs, as well as strains that modulated mouse adiposity and cecal metabolite concentrations, using feature selection algorithms and follow-up monocolonizations. This combinatorial approach enables a systems-level understanding of microbial contributions to human biology.
© 2014 Medical Xpress