Using computers to model our computer, the brain
The human brain is the most complex computer in existence. Understanding how it works has been a scientific endeavor for centuries. However, technology has only recently advanced to the point where we can really understand brain function down to the molecular level. By combining the newest advances in computer programming with biochemistry, Okinawa Institute of Science and Technology Graduate University's Computational Neuroscience Unit is helping to create the tools that will advance our understanding of the brain. Members of Professor Erik De Schutter's unit have developed a computational simulator called STEPS to help researchers understand neuronal signaling pathways at a molecular level. Since its first release in 2009, this simulator has been significantly enhanced. In a recent paper published in Frontiers in Neuroinformatics, researcher Dr. Weiliang Chen introduced two new toolkits he has developed to further extend STEPS functionality.
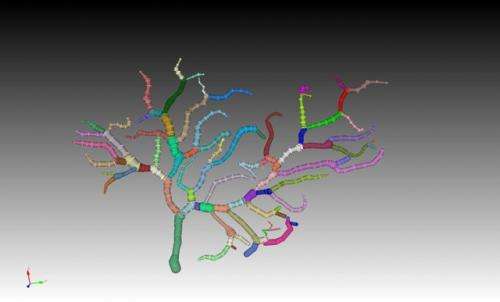
Computer modeling allows researchers to collect data and anticipate what may happen in a live brain, helping them test and develop hypotheses that can be used in conjunction with experiments in the lab. Computational neuroscience simulators, such as STEPS, rely on detailed reconstructions of the complex morphology, or shape, of the neurons. The reconstruction of neuronal morphology for simulation is difficult and time consuming, although recent advances in scientific software has somewhat simplified the process. The geometry preparation toolkit introduced in the paper simplifies recreation of neuronal morphology for the researcher. The second toolkit, the visualization toolkit, allows researchers to directly visualize STEPS simulations of those reconstructed morphologies in real time, both graphically and numerically, a capability that was not available in the STEPS program before.
In the paper, Chen demonstrates how the toolkits work in two different simulations. One example, using the geometry preparation toolkit, illustrates how the complex morphology of a neuron impacts the function of the cell. In this simulation of anomalous diffusion, you can see that the molecules get caught in the protrusions of the cell, the 'spiny' parts of what are called spiny dendrites, and diffuse slower than the ones in a smooth dendrite that does not have the protrusions. The other example, demonstrating the visualization toolkit, simulates what happens when an ion channel opens to release stored calcium within a neuron. Researchers are able to see the effects of the channels opening and the calcium being released both graphically and numerically. Both of these examples have been previously reported in laboratory experiments and simulated using computational modeling. However, the ability to easily recreate the shape of the cell, watch these events as they occur and manipulate conditions in real time is due to the new toolkits.
The entire STEPS project is part of a worldwide research initiative by the European Commission called the Human Brain Project. Ultimately, the goal is to integrate STEPS, which models neurons at a molecular level, with other computational models that look at the cellular and neural network levels that others are developing, to create a massive virtual human brain. Researchers will use this virtual brain to study anything from traumatic brain injury to the effect of a drug or a disease. The Computational Neuroscience Unit is one of only two laboratories in Japan that are part of the Human Brain Project.
As for Chen, he says, "I just want to see what's going on in the brain, and I want others to see it easily as well!" Asked whether or not the Human Brain Project will create that understanding, he says, "eventually we will be there, definitely. But you need to get the tools ready before that can happen." With ingenuity from researchers like him, it seems we are well on our way.
More information: Python-based geometry preparation and simulation visualization toolkits for STEPS. Weiliang Chen and Erik De Schutter. Front. Neuroinform., 11 April 2014 | DOI: 10.3389/fninf.2014.00037, journal.frontiersin.org/Journa … ninf.2014.00037/full