One route to malaria drug resistance found
Researchers have uncovered a way the malaria parasite becomes resistant to an investigational drug. The discovery, at Washington University School of Medicine in St. Louis, also is relevant for other infectious diseases including bacterial infections and tuberculosis.
The study appears July 24 in Nature Communications.
Many organisms, including the parasite that causes malaria, make a class of molecules called isoprenoids, which play multiple roles in keeping organisms healthy, whether plants, animals or bacteria. In malaria, the investigational drug fosmidomycin blocks isoprenoid synthesis, killing the parasite. But over time the drug often becomes less effective.
"In trials testing fosmidomycin, the malaria parasite returned in more than half the children by the end of the study," said senior author Audrey R. Odom, MD, PhD, assistant professor of pediatrics. "We wanted to know how the parasite is getting around the drug. How can it manage to live even though the drug is suppressing these compounds that are necessary for life?"
Fosmidomycin, an antibiotic, is being evaluated against malaria in phase 3 clinical trials in combination with other antimalarial drugs.
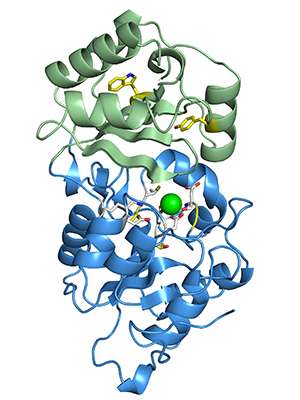
Using next-generation sequencing technology, the research team compared the genetics of malaria parasites that responded to the drug to the genetics of malaria parasites that were resistant to it. With this approach, Odom and her colleagues found mutations in a gene called PfHAD1. With dysfunctional PfHAD1, malaria is resistant to fosmidomycin.
"The PfHAD1 protein is completely unstudied," Odom said. "It's a member of a larger family of proteins, and there are almost no biological functions assigned to them."
In malaria parasites, Odom's team showed that the PfHAD1 protein normally slows down the synthesis of isoprenoids. In other words, when present, PfHAD1 is doing the same job as the drug, slowing isoprenoid manufacturing. Since isoprenoids are necessary for life, it's not clear why the organism would purposefully slow down isoprenoid production.
"We don't know why the protein puts the brakes on under normal conditions," Odom said. "Perhaps simply because it's an energetically expensive pathway. But loss of PfHAD1 releases the brakes, increasing the pathway's activity, so that even when the drug is there, it doesn't kill the cells."
Odom says isoprenoid synthesis is an attractive drug target not just for malaria but for tuberculosis and other bacterial infections because these organisms also rely on this same isoprenoid pathway. While people make isoprenoids, these vital compounds are manufactured entirely differently in animals compared with many infectious pathogens likely to cause disease.
Inhibiting isoprenoid manufacturing in malaria, bacteria or tuberculosis, for example, would in theory leave the human pathways safely alone. In people, perhaps the most well-known isoprenoid is cholesterol, with statin drugs famously inhibiting that manufacturing pathway.
Odom, who treats patients at St. Louis Children's Hospital, said she sees a handful of malaria cases each year, mostly in patients who have recently traveled to parts of the world where malaria is common. The parasite remains a massive global health problem, causing about 627,000 deaths in 2012 alone, according to the World Health Organization. Most deaths are in children under age 5.
Despite this public health burden, malaria is understudied in the lab because it is notoriously difficult to grow. It has a complex lifecycle that includes two-way transfers between mosquito and human and spans different forms in the human liver and red blood cells.
"The malaria parasite is difficult to work with in the lab; it's nearly impossible to replicate the lifecycle," Odom said. "That's why it was so exciting to be able to do this kind of study in malaria, rather than in a typical model organism like yeast. This genetic study would not have been possible even five years ago because the gene sequencing technology was not there."