Pitt drug discovery researchers receive grant to build 3-D liver model
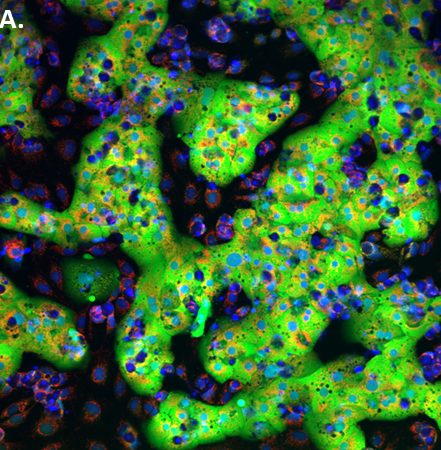
With a new $5.8 million, three-year award from the National Institutes of Health (NIH), researchers at the University of Pittsburgh School of Medicine will further develop a state-of-the-art, microfluidic 3D model system that mimics structure and function of the liver to better predict organ physiology, assess drug toxicity and build disease models.
The funding supports the next phase of the NIH's Tissue Chip for Drug Screening program, which aims to improve ways of predicting drug safety and effectiveness. Researchers from 11 institutions will collaborate over three years to refine existing 3-D human tissue chips and combine them into an integrated system that can mimic the complex functions of the human body.
"We are very enthusiastic about the potential of these microphysiology systems to serve as powerful platforms for studying human diseases and identifying human toxic liabilities," said the Pitt project's principal investigator D. Lansing Taylor, Ph.D., Allegheny Foundation Professor of Computational and Systems Biology, Pitt School of Medicine, and director, University of Pittsburgh Drug Discovery Institute.
"The development of tissue chips is a remarkable marriage of biology and engineering, and has the potential to transform preclinical testing of candidate treatments, providing valuable tools for biomedical research," said NIH Director Francis S. Collins, M.D., Ph.D.
The Pitt research team, along with additional collaborators, is creating models of the functional unit of the liver, called the acinus, using human liver cells and eventually liver cells derived from precursor cells known as induced pluripotent stem cells, as well as three additional cell types. The liver platform includes microfluidic devices, human cells, engineered matrix materials, fluorescence-based biosensors for real-time physiological read-outs, and biochemical and mass spectrometry measurements to determine acute and chronic toxicity effects. They also will build a "microphysiology database" to manage, analyze and model the data collected from the liver constructs.
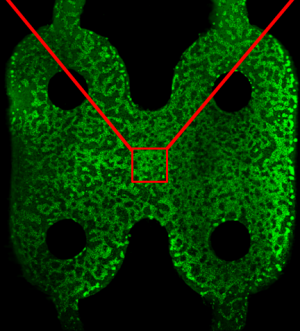
With such a platform, biomedical scientists will be able to test treatment efficacy in conditions such as non-alcoholic fatty liver disease, liver cancer and breast cancer that has spread to the liver, as well as liver damage including immune-mediated toxicology and fibrosis. Also, a team of institutions and investigators has been assembled to integrate the liver, kidney and gut models to recapitulate the organ system that is central to drug absorption and metabolism.
The integrated platform will involve the creation of a universal medium, the development of the proper "scaling" of the interacting organ constructs, physiologically relevant flow, incorporation of a micro-formulator to add factors from missing organs and micro-analyzers for monitoring parameters such as pH and oxygen.
Fifteen NIH Institutes and Centers are involved in the coordination of the tissue chip program. Current funding is being provided by the National Center for Advancing Translational Sciences, the National Institute for Biomedical Imaging and Bioengineering, the National Cancer Institute, Eunice Kennedy Shriver National Institute of Child Health and Human Development, National Institute of Environmental Health Sciences, NIH Common Fund and NIH Office of Research on Women's Health.
Collaborators include Martin Yarmush, M.D., Ph.D., of Massachusetts General Hospital; John Wikswo, Ph.D., of Vanderbilt University; Jonathan Himmelfarb, M.D., of the University of Washington; Mark Donowitz, M.D., of Johns Hopkins University; and Mary Estes, Ph.D., of Baylor University.