Scientists describe the function of an enzyme critical to male fertility
Researchers are one step closer to unraveling the extraordinarily complex series of processes that leads to an event crucial to human reproduction: the creation of sperm.
In a study published in the journal Genes and Development, University of Pennsylvania researchers have filled in details of how an enzyme, through interactions with a network of nearly two dozen other genes, protects the integrity of the germ line by giving rise to a class of RNA molecules that are essential to sperm development.
The enzyme of interest, the RNA helicase MOV10L1, is likely the source of mutations that cause some cases of male infertility and could one day serve as a target for a form of reversible male contraception.
The work was jointly led by Jeremy Wang, a professor of developmental biology in the School of Veterinary Medicine, and Zissimos Mourelatos, a professor of pathology and laboratory medicine in Penn's Perelman School of Medicine. They collaborated with co-lead authors Anastassios Vourekas and Manolis Maragkakis of Penn Medicine, Qi Fu of Penn Vet and Ke Zheng of Nanjing Medical University. Co-authors included Penn Medicine's Panagiotis Alexiou, Nanjing Medical University's Jing Ma and the European Molecular Biology Laboratory's Ramesh S. Pillai.
From earlier work, the group knew that MOV10L1 was required to produce a class of small RNA molecules called piwi-interacting RNAs, or piRNAs. piRNAs are essential to germ-line development in males.
Wang, Mourelatos and colleagues wanted to learn more about the mechanism by which MOV10L1 generated piRNAs.
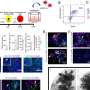
Because MOV10L1 is believed to be an RNA helicase, or an enzyme that can unravel RNA molecules, they first performed a screening assay, known as a HITS-CLIP, to catalog the RNA transcripts to which MOV10L1 bound, looking in the testes of normal adult mice. The researchers found that almost all the RNAs that MOV10L1 was binding were precursors of piRNAs.
"This suggests that MOV10L1 is very specific in what it binds," Wang said. "It's pretty much only interacting with this class of RNA precursors."
Looking more closely at the areas where MOV10L1 bound to RNAs, they found a nucleotide preference; the bound RNA transcripts were enriched for guanine (G) nucleotides. This finding led the researchers to suspect that the enzyme was preferentially binding RNA where the transcript bent into arrangements called G-quadruplexes, or G4s. G4s are very stable arrangements in which RNA molecules fold in such a way that they create "stacks" of interconnected Gs that are difficult to break.
The Penn-led team also noted that G4s and other RNA secondary structures predominated in the tail end of mature piRNAs, a pattern that was present in human, monkey, mouse and rat genomes. The fact that this pattern was conserved across species provides strong evidence that it plays a crucial role in piRNA processing in these organisms.
This find led the researchers to a hypothesis: that these G4 structures, because they are so stable and difficult to break up, serve as "road bumps" at which MOV10L1 and its accompanying proteins pause during the processing of long RNA transcripts.
To test whether the helicase activity of MOV10L1 was responsible for processing piRNAs, the researchers generated a mutated form of the enzyme in which a particular amino acid was changed in the region believed to be essential for its enzymatic activity. In test tube-based experiments, they found that while the normal enzyme was able to unwind RNA structures, the mutated form could not. As they anticipated, when mice with this mutant MOV10L1 were bred, the males were sterile and resembled mice that lacked MOV10L1 altogether.
"We think that this MOV10L1 complex is like a car, moving along a single-stranded RNA," Wang said. "When it hits this G4 secondary structure, the complex cannot move any more so it pauses, which then allows another enzyme to cut in that location and release part of the transcript for additional processing. MOV10L1 then uses its helicase activity to untie the knots of the G-quadruplex, straightening it out and resolving the structure. This process happens repeatedly along the RNA precursor."
"This complex appears to present the long strand of RNA to the nuclease and initiate the first cut, which is important for piRNA biogenesis," Mourelatos said.
These chopped bits of RNA give rise to the mature piRNAs. piRNAs preserve the integrity of the germ line during sperm production by blocking the effects of retrotransposons, which could otherwise introduce damaging mutations.
Both Wang's and Mourelatos' labs plan to continue probing the MOV10L1 complex to elucidate more steps involved in the production of piRNA. Mourelatos is interested in the connection between piRNA biogenesis and mitochondria, where the RNAs are processed and where organisms produce most of the energy required to live.
"This leads us to wonder whether there is a metabolic connection that we don't understand," Mourelatos said.
Wang and colleagues also intend to search for ways to block this pathway.
"If we block the interactions between MOV10L1 and its associated proteins, we may be able to disrupt this complex," Wang said. "And if you disrupt it, you would disrupt piRNA production, and that could have a contraceptive effect."
Wang noted that blocking the production of piRNA at this stage could result in potentially reversible contraception, preventing sperm from being formed.