Scientists pinpoint mutations responsible for ineffective 2014-2015 flu vaccine
Viruses like influenza have the ability to mutate over time, and given that the flu vaccines administered during the 2014-2015 season were largely ineffective at preventing the spread of the flu, it appears the virus that recently circulated had taken on mutations not accounted for when last year's vaccine was developed.
Now, researchers at The Wistar Institute identified specific mutations that influenza recently acquired to escape the current vaccine design.
Scott Hensley, Ph.D., Assistant Professor at The Wistar Institute, led a study published online by the journal Cell Reports.
Each year, seasonal influenza sickens 3 to 5 million people worldwide and is responsible for up to half a million deaths.
When the flu vaccine is designed each year, it is meant to protect against specific viral strains that are expected to circulate. The vaccine elicits antibodies that respond to those specific strains when a person gets infected with the virus. If the antibodies match the strain, then a person should be protected from the flu virus in a given year.
However, specific antibodies that are elicited by vaccination can be ineffective when the virus acquires mutations at sites where the antibodies are supposed to bind, which are called antigenic sites. In a phenomenon known as antigenic drift, mutations accumulate in these sites, thus creating new viruses for which previously effective antibodies can no longer fight against. This leads to vaccines that are less able to protect against the virus. According to some estimates, only about one in four people who received last year's seasonal flu vaccine were protected against the virus.
"Our studies show that flu viruses recently acquired mutations in critical regions that are recognized by our immune system," Hensley said. "These new mutations likely contributed to the ineffectiveness of flu vaccines during the 2014-2015 flu season."
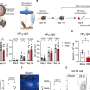
Researchers in Hensley's lab identified 10 different viral mutations that circulated from 2014-2015 and differed from the viral strain that was used to create the vaccine. They then examined sera - the clear part of the blood that contains viral antibodies -from humans as well as ferrets and sheep exposed to the current H3N2 vaccine strain. This was done because ferrets and sheep can be infected with and respond similarly to the same influenza strains that affect humans. They found that mutations in a specific region of H3N2 viruses significantly decrease the effectiveness of these antibodies.
Researchers in Hensley's lab began investigating antigenic drift of the 2014-2015 seasonal influenza viruses in December of 2014 while the virus was still circulating. The mutations were studied using reverse engineering and allowed them to modify the ineffective H3N2 vaccine strain so that it matched the H3N2 strain with mutations caused by antigenic drift.
"I have a really talented team of researchers who are very good at quickly dissecting the specificity of flu antibody responses," Hensley said.
"We identified mutations that were common in flu isolates in December of 2014 and we engineered viruses that allowed us to characterize these mutations the following month," said Benjamin Chambers, a graduate student in the Hensley laboratory.
The World Health Organization (WHO) has recommended that the seasonal flu vaccine be appropriately updated for the 2015-2016 flu season to account for this recent case of antigenic drift. The authors note that their data strongly supports the WHO's recommendation to update the flu vaccine for future seasonal influenza infections, and they are currently completing studies to determine if antibodies elicited by the new vaccine strain recognize the types of H3N2 strains that might circulate next year.
More information: Cell Reports, Chambers et al.: "Identification of Hemagglutinin Residues Responsible for H3N2 Antigenic Drift during the 2014-2015 Influenza Season" dx.doi.org/10.1016/j.celrep.2015.06.005