Regenerating damaged cardiac muscle
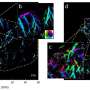
To mend a broken heart—that is, to regenerate a damaged cardiac muscle—it helps to know how hearts are built. "How does one stem cell, which has no specific identity, develop into multiple cell types that organize into this beautiful three-dimensional structure?" asks Laurie Boyer, associate professor in the MIT Department of Biology, who tackles that problem by investigating regulatory elements that switch genes on and off at the right time and place during development. Faulty regulation can lead to congenital heart defects, which are the greatest cause of infant morbidity and mortality.
Heart cells must be "super-strong" to withstand continuous pumping throughout life, she says. Yet they stop dividing shortly after birth, losing the ability to repair or replace themselves if damaged by a heart attack. In contrast, skin or intestinal cells can continuously regenerate.
Boyer began her career at a time when genome sequencing started to reveal that gene regulation was more complex than anticipated. She focused on learning how networks of genes are regulated during heart development, because by learning how these genes are turned on or off, she hoped to understand congenital heart disease. In a 2013 paper in Cell, she characterized a novel gene belonging to a poorly understood class of molecules called long non-coding RNAs (lncRNAs). The gene, which she named Braveheart, helps specify which early cells will develop as heart cells. Braveheart was the first lncRNA implicated in heart development, and its discovery revealed an entirely new, previously unsuspected regulatory layer.
In a series of papers from 2012 to 2015, she describes her investigations of these processes in mice. Cardiac cells in mice continue actively dividing during their first week of life, and then stop after a discrete interval. During that interval, if the heart is injured, cardiac cells revert to a more immature state. They begin dividing again, and those new cells mature into functional cardiac muscle cells.
"We realized we could exploit this short developmental window to identify the molecular roadblocks to regeneration. What changes in gene regulation happen in the injury response?" Some of those changes involve regulatory elements called enhancers: short DNA sequences that bind to transcription factors (which are proteins that actually turn genes on and off) and that act very specifically in different tissues at discrete developmental stages.
"Now we are asking: How can we turn back the developmental clock for mature cardiac cells? We'd like to manipulate the gene regulatory circuitry so we can reprogram human cardiac cells to repair themselves after injury." She is focused on transcription factors because as proteins, they are easier to target. Preliminary results point to factors not previously known to play a role in heart development and disease.
Boyer is excited to gain new knowledge about the heart, especially if it might one day help develop cures for heart defects and disease. She had no idea where her interest in stem cell development would lead. "Success is different for everyone," she advises young scientists, "and its measure for me is how I overcome obstacles to follow my passion."
This story is republished courtesy of MIT News (web.mit.edu/newsoffice/), a popular site that covers news about MIT research, innovation and teaching.