October 2, 2015 report
Technique identifies prenatal trisomy, cancer type, and transplant rejection using methylation sequences of plasma DNA

Researchers from China were able to detect liver cancer and lymphoma in cancer patients, genetic abnormalities from the placenta in pregnant women, and donor DNA in post-transplant patients by analyzing the methylation sequence in plasma DNA. Using known tissue methylation profiles, they were able to identify the tissue origins of circulating DNA, making this a robust tool for screening, cancer detection, and therapy monitoring. This collaborative effort by researchers from several institutions is published in the Proceedings of the National Academy of Sciences.
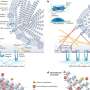
Epigenetics have shown that DNA will often have methyl groups attached to the 5-carbon on cytosine. Certain tissues, and certain cell types within the tissues, have a distinct methylation pattern in its DNA, allowing scientists to create a methylome database of methylation patterns for various tissues in the body. While scientists have been able to search plasma for DNA from a single tissue type, such as placental or liver tissue, Sun, et al. used genome-wide bisulfite sequencing to elucidate the methylation pattern of plasma DNA and compared it to the methylome database to discern the percent composition, or tissue map, of the plasma DNA. It provides what the authors call a "bird's eye view" of the plasma, showing the origin of the circulating DNA.
To test their technique, Sun, et al. identified 5,820 methylation markers that they would use to then run through a deconvolution algorithm to identify the tissue or cell type based on its methylation pattern. They then tested their algorithm using DNA of various compositions of buffy coat DNA, placenta DNA, and liver DNA.
Once the procedure was tested, it was time to test whether the procedure could accurately identify compositional changes. The first trial was with fifteen pregnant women, five in each of the first, second, and third trimester. Their plasma should have more DNA from the placenta compared to non-pregnant women. They found that white blood cells were the largest contributor to the plasma DNA, something that had only been observed in bone marrow transplants, but had not been confirmed in other cases. Secondly, they also found that 12.1% to 41.0% of its DNA was from the placenta. The percentage of placenta DNA was significantly less in the control group. The percentages were verified using paternally inherited fetal SNP alleles.
The second test involved people who had received a transplant. Development of graft-versus-host disease is a common problem in allogeneic transplants. Donor DNA has been found in transplant patient's blood plasma. Four liver transplant recipients and three bone marrow transplant recipients' plasma DNA were analyzed and compared to donor SNP alleles in the case of the liver patients, and white blood cells in the case of the bone marrow patients. They were able to detect donor DNA, and found a strong correlation between the results from analyzing the methylation pattern and looking at SNP alleles or white blood cells.
Sun, et al. then determined if their technique would work for detecting cancer cells in twenty-nine subjects with hepatocellular carcinoma (HCC) compared to subjects without cancer. The plasma should have a higher percentage of DNA from the cancerous tissue compared to subjects without cancer. Their algorithm showed that the median contribution by the liver to the plasma in the HCC subjects was 24.0% compared to 10.7% in the control subjects. To check their methods, they also compared their results to tumor DNA concentration using a previously reported method.
Given these experimental results, the authors then tested whether they could identify the tissue of origin based on plasma copy number aberrations found in plasma DNA. If additional copies are made, then there should be a greater percentage of DNA from that tissue compared to a genomic region that does not have a copy number aberration. The same reasoning can be used for copy number deletions in which there would be less DNA from a particular tissue type compared to other genomic regions.
They showed that when looking at the plasma of women with fetuses containing an extra chromosome (trisomy 21), the amount of placental tissue was higher in methylation markers in chromosome 21 compared to markers in other chromosomes. The other tissues (liver, neutrophils, and lymphocytes) did not show and increase in copy number.
They then looked at amplifications or deletions in plasma DNA of subjects with HCC. These were compared to plasma DNA that did not show amplifications or deletions. They were able to determine that seven of the HCC subjects showed copy number aberrations and that these portions contained more DNA from the liver.
During this study, the researchers identified a 37-year-old pregnant woman whose lymphoma had returned. She was first diagnosed in 2011 and underwent chemo therapy. There was no more detectable lymphoma after chemo and in subsequent check-ups. Then, in March 2014 at eleven weeks pregnancy, blood samples collected for noninvasive prenatal testing showed that the maternal plasma had some abnormalities. Lymph node biopsy confirmed that the lymphoma had returned. Comparison of the plasma DNA to the lymph node biopsy showed similar copy number aberrations. However, ten weeks after chemo therapy, plasma DNA showed no copy number aberrations. Furthermore, they found that the origin of the copy number aberrations were from B lymphocytes, but not T lymphocytes.
With additional research, this technique may be an excellent noninvasive way to assess several factors in a patient's health. Indeed, the authors point out that this work has created a bridge between molecular diagnostics and the traditional more organ-based medical practices. The last study demonstrated the benefit of being able to identify the origin of the DNA in plasma since the test was intended to identify chromosomal abnormalities in the fetus, but the abnormalities had actually originated from the mother. Lymph node biopsy confirmed the re-occurrence of her cancer.
More information: "Plasma DNA tissue mapping by genome-wide methylation sequencing for noninvasive prenatal, cancer, and transplantation assessments" PNAS, DOI: 10.1073/pnas.1508736112
Abstract
Plasma consists of DNA released from multiple tissues within the body. Using genome-wide bisulfite sequencing of plasma DNA and deconvolution of the sequencing data with reference to methylation profiles of different tissues, we developed a general approach for studying the major tissue contributors to the circulating DNA pool. We tested this method in pregnant women, patients with hepatocellular carcinoma, and subjects following bone marrow and liver transplantation. In most subjects, white blood cells were the predominant contributors to the circulating DNA pool. The placental contributions in the plasma of pregnant women correlated with the proportional contributions as revealed by fetal-specific genetic markers. The graft-derived contributions to the plasma in the transplant recipients correlated with those determined using donor-specific genetic markers. Patients with hepatocellular carcinoma showed elevated plasma DNA contributions from the liver, which correlated with measurements made using tumor-associated copy number aberrations. In hepatocellular carcinoma patients and in pregnant women exhibiting copy number aberrations in plasma, comparison of methylation deconvolution results using genomic regions with different copy number status pinpointed the tissue type responsible for the aberrations. In a pregnant woman diagnosed as having follicular lymphoma during pregnancy, methylation deconvolution indicated a grossly elevated contribution from B cells into the plasma DNA pool and localized B cells as the origin of the copy number aberrations observed in plasma. This method may serve as a powerful tool for assessing a wide range of physiological and pathological conditions based on the identification of perturbed proportional contributions of different tissues into plasma.
© 2015 Medical Xpress