Researchers elucidate network of genes that control when puberty begins

In expanding our knowledge of how the brain controls the process of sexual development, researchers at Oregon Healthy & Science University and the University of Pittsburgh have identified for the first time members of an elaborate superfamily of genes that regulate the timing of puberty in highly evolved nonhuman primates. The Zinc finger, or ZNF, gene family comprises approximately 800 individual genes.
A handful of genes in this network, operating within the neuroendocrine brain, serve as a 'neurobiological brake' that delay until the end of childhood the activation of hypothalamic genes responsible for launching puberty, thereby preventing the premature awakening of the process. The paper was published today in the journal Nature Communications.
The paper demonstrates that the ZNF gene family encodes repressors—proteins that hold in check the activity of genes—to suppress the launch of puberty. The researchers' fresh insights better position scientists to decipher whether environmental factors push the start of puberty to younger ages. Early puberty is associated with an increased incidence of ovarian, uterine and breast cancer as well as an increased incidence of cardiovascular disease and metabolic diseases.
"Deepening our understanding of how the brain controls the initiation of puberty will allow us to understand why girls are initiating puberty at much earlier ages. This knowledge may help build a foundation for developing new avenues to treat precocious puberty," said Alejandro Lomniczi, Ph.D., lead researcher on the study and assistant scientist in neuroscience at the Oregon National Primate Research Center at OHSU. "Our suspicion is that chemical substances contained in man-made products and other environmental factors, such as nutrition, may accelerate reproductive development by epigenetically antagonizing gene repressors such as ZNFs".
ZNFs exert their inhibitory effect by setting in motion mechanisms that modify gene activity without changing the sequence of DNA. Because of this, the ZNFs are considered to act "epigenetically," that means by conveying to genes information from the environment without changing the genetic code itself.
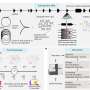
The researchers found that the abundance of the messenger RNAs encoding GATAD1 and ZNF573, along with that of five other ZNFs, decreases during the juvenile-pubertal transition in nonhuman primates, when the brake on the hypothalamic drive to the pituitary-gonadal axis is released. ZNFs promote the loss of a DNA-associated protein known as 'histone 3 dimethylated at lysine 4' (H3K4me2) from the controlling region of genes that facilitate puberty. H3K4me2 is normally associated with gene activation. Using a gene therapy paradigm and a rodent model, the researchers selectively increased the abundance of GATAD1 or ZNF573 in the hypothalamus of prepubertal female rats and observed that puberty was delayed in these animals. Altogether their findings suggest that as the production of ZNFs decreases in the hypothalamus during late prepubertal development, the "brake" keeping puberty in check is released and sexual maturity begins.
More information: Nature Communications, dx.doi.org/10.1038/NCOMMS10195