New understanding of TB could lead to personalised treatments
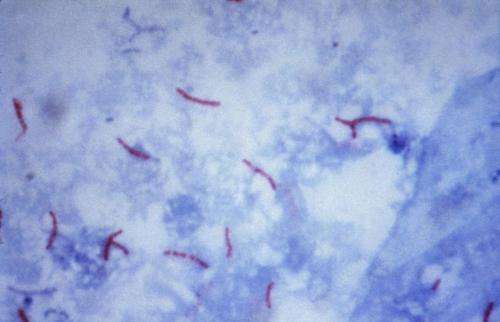
Tuberculosis (TB) used to kill one in seven people around the world before the advent of antibiotics. However the increasing prevalence of multi-drug resistant TB means that doctors are running out of options in trying to combat this bacterium.
In addition, globally TB currently ranks alongside HIV as the leading cause of death worldwide, killing 1.5 million in 2014.
Now a team from Trinity College Dublin and St James's Hospital have made a breakthrough in terms of how our immune system responds to TB, which opens up the possibility to design more effective vaccines and personalised therapies.
Dr Clíona Ní Cheallaigh, lead author on this study, which has just been published in the journal Immunity, was struck by how different people all seemed to respond differently to the tuberculosis bacteria.
For example, thinking back to the Dublin tenements of a hundred years ago, crowded conditions meant that TB was easily spread among communities yet not everyone who became infected got sick. This is unlike most infectious diseases.
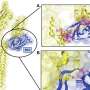
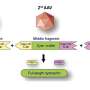
Dr Ní Cheallaigh, along with colleagues and senior authors on the paper Professor in Respiratory Medicine, Joe Keane, St James's, and Professor in Immunology Ed Lavelle, Trinity Biomed Sciences Institute (TBSI), focused on the protein Mal and how it impacts on responses to TB.
Mal was of interest because about 25% of Europeans have a different form of the protein and these tended to be more susceptible to getting sick with TB but up to now it was not clear why.
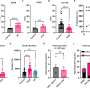
What this research, funded by the Health Research Board, the Royal City of Dublin Trust and Science Foundation Ireland, shows for the first time is that Mal is involved in cell signalling in response to Interferon Gamma, a master chemical in our immune response to illness. Producing Interferon Gamma is like putting a fire-lighter on a fire – it sets off a strong immune response when you have an infection.
Dr Ní Cheallaigh commented on the findings: "Having this different form of Mal affects how intensely you respond to Interferon Gamma – if you've one form you have a big response, if you have another form you have a dampened down response. We've discovered a whole new function for this protein Mal."
HRB Clinician Scientist Prof Joe Keane further added: "In St James's Hospital, we treat drug-resistant TB patients who need novel immune treatments, like Interferon, which can be optimised because of this research."
The finding is significant because it means that patients with TB, especially difficult to treat cases, can benefit from more targeted treatments personalised to their specific immune system.
Prof Ed Lavelle stressed that this discovery also has implications in the treatment of other illnesses where Interferon Gamma is involved. These include a number of infectious diseases and cancer and the form of Mal we have may also influence how we respond to vaccines.
More information: Clíona Ní Cheallaigh et al. A Common Variant in the Adaptor Mal Regulates Interferon Gamma Signaling, Immunity (2016). DOI: 10.1016/j.immuni.2016.01.019