Computer-assisted approaches as decision support systems serving to combat the Zika virus
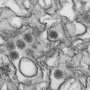
Global climate change, international travel, and ineffective vector control programs are aiding the emergence of infectious diseases globally. The currently expanding Zika virus (ZIKV) epidemic is one such problem. The rapid expansion of this disease to epidemic proportions in South America in 2015-16 has led the World Health Organization to declare ZIKV a public health emergency on February 1, 2016. Two main reasons behind this are suspected association of the virus to cases of microcephaly in children born of ZIKV-infected mothers and Guillain- Barré syndrome (GBS)—an autoimmune disease that may occasionally lead to a fatal form of paralysis.
ZIKV is transmitted primarily by two mosquito vectors: 1) Aedes aegypti, prevalent mostly in tropical climates, and 2) Aedes albopictus which ranges in the Americas up to the Great Lakes. Mass gatherings like carnivals and Olympics, from where infected visitors may carry the virus to their countries around the globe, have the potential of expanding ZIKV infection into pandemics. No drug is known to treat ZIKV infection; neither do we have any vaccine which can prevent the spread of the virus. While scientists and global health authorities are trying to cope with the situation, computer-assisted approaches itemized below may help as decision support systems:
1) COMPUTER-ASSISTED DRUG DESIGN
Once some bioassay is developed, computer-assisted methods can help to select manageable subsets from large chemical libraries for testing in high throughput screening (HTS) protocols. Pharmacophores derived from active molecules can also be used to screen libraries for ZIKV drug discovery.
2) COMPUTER-AIDED VACCINE DESIGN
Computational approaches to peptide vaccine design have been developed by us. Because peptides are cheaply available from commercial vendors, lab testing of candidate peptides as potential ZIKV vaccine can be pursued within a reasonable time frame and modest funding.
3) MATHEMATICAL APPROACHES FOR THE CHARACTERIZATION OF EMERGING ZIKV STRAINS
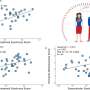
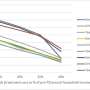
Alignment-free mathematical descriptors for characterization of DNA/ RNA sequences of pathogens like H5N1 and H5N2 pandemic bird flu have proved effective for comparative studies. A battery of mathematical sequence descriptors may provide us a quantitative view of the sequences, in whole or in part as required and may aid in the surveillance of emerging strains.
The topic is discussed in detail in the editorial article, Computer-assisted approaches as decision support systems in the overall strategy of combating emerging diseases: Some comments regarding drug design, vaccinomics, and genomic surveillance of the Zika virus, published in the journal, Current Computer-Aided Drug Design.
More information: Subhash Basak et al. Editorial: Computer-Assisted Approaches as Decision Support Systems in the Overall Strategy of Combating Emerging Diseases: Some Comments Regarding Drug Design, Vaccinomics, and Genomic Surveillance of the Zika Virus, Current Computer Aided-Drug Design (2016). DOI: 10.2174/1573409912999160315115502