Detecting brain tumors' warped metabolism
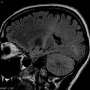
When a brain tumor is suspected because of symptoms such as headaches or other problems, its presence is usually confirmed by anatomical imaging such as CT or MRI. But through imaging, doctors often can't say much about the tumor – molecularly – besides "something's in there." Surgery and a biopsy are necessary to get a glimpse of the cancer cells themselves.
That changed with the recent discovery that some brain tumors have mutations that alter cancer cell metabolism, producing a distinctive chemical or "oncometabolite" (D-2-hydroxyglutarate) that can be seen via metabolic imaging non-invasively. The levels of this chemical are much lower in normal brain cells.
Erwin Van Meir, PhD and his colleagues at Winship Cancer Institute of Emory University have now shown that D-2-hydroxyglutarate can also be detected in cerebrospinal fluid (CSF), the liquid that bathes and protects the brain. Scientists analyze the cerebrospinal fluid using mass spectrometry, which is many times more sensitive than imaging techniques.
The results were published Thursday, June 23 in Clinical Cancer Research.
The authors describe their mass spectrometry-based approach as complementary to the detection of D-2-hydroxyglutarate by imaging. While it is somewhat more invasive to draw cerebrospinal fluid, it has advantages over imaging and could be used repeatedly to track the effectiveness of anticancer treatment.
"These findings further demonstrate the potential of CSF as a source for brain tumor biomarkers that could be useful for diagnosis, prognosis and monitoring of treatment efficacy, thus providing a window on brain tumor biology without the need for repeated surgical tumor sampling," Van Meir says.
Van Meir is a professor of neurosurgery and hematology and medical oncology at Emory University School of Medicine, leader of the Cancer Cell Biology program at the Winship Cancer Institute, and founding director of Emory's graduate program in Cancer Biology.
The mutations that cause the alterations in metabolism are in the genes for isocitrate dehydrogenase (IDH1 or IDH2), and are found in more than 75 percent of low- and intermediate-grade gliomas, and in 10 percent of a more aggressive form of brain cancer, glioblastoma. The D-2-hydroxyglutarate appears to warp how the cancer cell maintains genes as activated or repressed.
Patients with tumors bearing these mutations have improved overall survival compared with similar brain tumors without them, but they are less responsive to standard radiation and chemotherapy treatment. Development of drugs, by a private company, targeting the mutated IDH1 and 2 enzymes is underway.
Former postdoc Juliya Kalinina, PhD, now a senior scientist at Merck, and senior research specialist Jun Ahn are the co-first authors of the paper, with the collaboration of radiologist Hui Mao, PhD and geneticist Miao He, PhD.
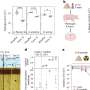
The mass spectrometry approach was validated by testing CSF samples from three cohorts of cancer patients (84 patients total), obtained from Penn State, University of Massachusetts Medical School, University of Mainz, UCSD and Utah's Huntsman Cancer Center. The average level of D-2-hydroxyglutarate in CSF was 17 times higher in patients with tumors bearing IDH mutations than in those without them.
More information: J. Kalinina et al. Selective detection of the D-enantiomer of 2-Hydroxyglutarate in the CSF of glioma patients with mutated Isocitrate Dehydrogenase, Clinical Cancer Research (2016). DOI: 10.1158/1078-0432.CCR-15-2965