Researchers create genetic blueprint of cunning tuberculosis bacteria
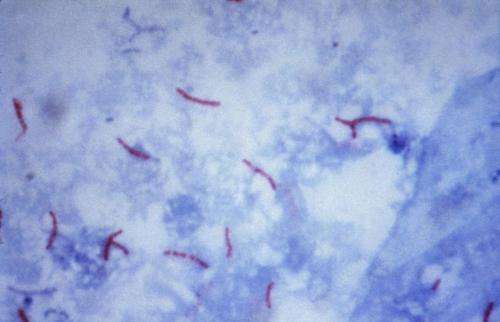
The rise in multi-drug resistant (MDR) and extremely drug resistant (XDR) strains of Mycobacterium tuberculosis (MTB) is becoming a major cause of global health concern for treating tuberculosis, which affects a third of the global population. In fact, the number of worldwide deaths caused by tuberculosis has surpassed HIV/AIDS, and there is greater sense of urgency than ever before to find effective drug cocktails to outsmart MTB.
In a landmark study to be published on June 6, 2016, in Nature Microbiology, researchers at Institute for Systems Biology and the Center for Infectious Disease Research in Seattle demonstrated a systems biology approach that has the potential to rationally predict combinations of drugs that will disrupt tolerance networks in MTB, making it even most susceptible to antibiotic therapy.
"The incredibly large number of possible drug combinations taken together with the difficulty of growing MTB in the laboratory make discovery of effective combination therapy extremely challenging. We hope that our systems-based strategy will accelerate TB drug discovery by helping researchers prioritize combinations that are more likely to be effective," said Dr. Nitin Baliga, of Institute for Systems Biology and the senior author on the paper.
The success of MTB is largely due to its ability to alter gene expression to counteract host defense and anti-tubercular drug treatment. An extended period of tolerance gives MTB a window of opportunity to mutate and evolve longer term resistance. Previously, researchers in the Baliga lab at ISB and in the Sherman lab at CIDR published a genome-wide regulatory network model that could predict how MTB senses and responds to changes in its environment, including anti-tubercular treatment.
In this study, they used the network model to understand how MTB tolerates killing by the drug bedaquiline, which in 2012 was the first drug in 40 years to be approved by the FDA. They used this network-enabled knowledge to find a second drug (pretomanid) to counteract tolerance against bedaquiline. They went on to demonstrate that making the tolerance network hyperactive abolished the effectiveness of the combination therapy, confirming the mechanism of combined action of bedaquiline and pretomanid. The success of this systems biology-based method to find drug combinations has the potential to revolutionize and rapidly accelerate efforts towards TB drug discovery.
More information: Network analysis identifies Rv0324 and Rv0880 as regulators of bedaquiline tolerance in Mycobacterium tuberculosis, Nature Microbiology, DOI: 10.1038/nmicrobiol.2016.78