Single gene mutation causes severely debilitating disease of brain blood vessels
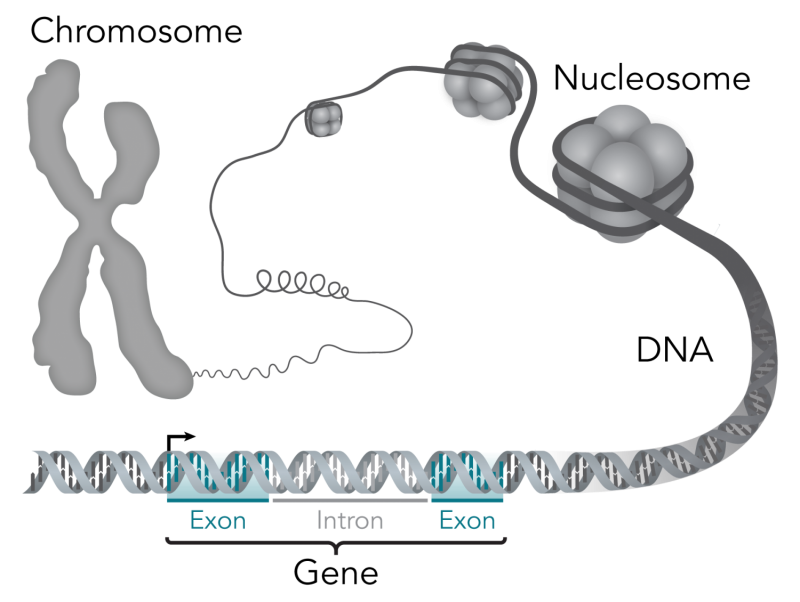
The cause of a disease that targets blood vessels in the brain - leading to multiple debilitating symptoms and, often, to early death - has been tracked to a single mutated gene, opening up the immediate possibility of improved patient care through genetic testing, and of future treatments.
The research, conducted over 12 years, was led by the teams of Professor Yanick Crow and Dr Ray O'Keefe from The University of Manchester's Division of Evolution and Genomic Sciences, and included contributions from over 60 academics and scientists from around the world. It is published today (29 August 2016) in the journal Nature Genetics.
This discovery involved the identification of mutations in SNORD118 - a small nucleolar RNA (snoRNA) gene - to cause leukoencephalopathy with calcification and cysts (LCC). LCC is characterised by progressive white matter degeneration in the brain. The disease can present in patients at any stage of life.
Dr Emma Jenkinson, Research Associate at The University of Manchester's Division of Evolution and Genomic Sciences, was lead laboratory researcher on this project. She said that the 12-year study unlocked greater understanding of the development of genetic disorders caused by mutations in non-coding regions of the genome: "The identification of SNORD118 as the instigator of a progressive and frequently fatal brain disease is a very significant step forward in understanding the role of non-coding RNAs in the development of certain diseases.
"Although LCC is relatively rare, by understanding the mechanism of this disease we may be able to extrapolate similar research into other diseases in the future - with the ultimate long term aim being the discovery of new treatments for patients."
The team collected biological samples and clinical data from 33 families affected by LCC, and used mapping techniques combined with next generation genome sequencing to identify the mutated gene. That then provided the team with evidence of 36 rare sequence variants, of which several were shown experimentally to cause a dysfunction of the snoRNA.
More information: 'Mutations in SNORD118 cause the cerebral microangiopathy leukoencephalopathy with calcifications and cysts' is published on 29 August in Nature Genetics, DOI: 10.1038/ng.3661