Researchers identify key elements of cellular response to proteasome dysfunction
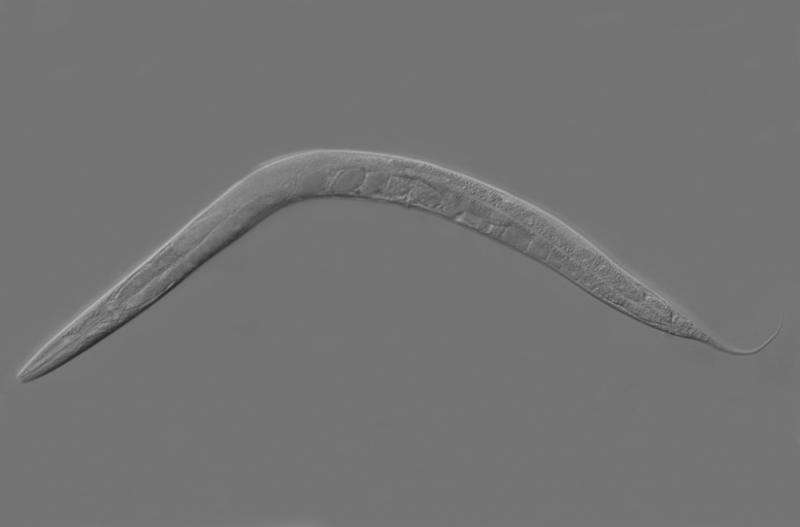
Maintaining appropriate levels of proteins within cells largely relies on a cellular component called the proteasome, which degrades unneeded or defective proteins to recycle the components for the eventual assembly of new proteins. Deficient proteasome function can lead to a buildup of unneeded and potentially toxic proteins, so cells usually respond to proteasome dysfunction by increasing production of its component parts. Now two Massachusetts General Hospital (MGH) investigators have identified key molecules in the pathway by which cells in the C. elegans roundworm sense proteasome dysfunction, findings that may have application to treatment of several human diseases.
"Proteasome inhibitors are currently being used to treat some cancers, including multiple myeloma, so future drugs targeting this pathway could enhance their activity," says Gary Ruvkun, PhD, of the MGH Department of Molecular Biology, corresponding author of the report in the journal eLife.
Lead author Nicolas Lehrbach, PhD, a postdoctoral fellow in Ruvkun's laboratory, adds, "Boosting proteasome activity might help treat some neurodegenerative disorders - including Alzheimer's and Parkinson's disease, both of which are characterized by abnormal protein deposits in the brain that exceed the ability of the proteasome to respond."
While it has been known that a transcription factor called SKN-1 was essential for the response to proteasome dysfunction in C. elegans, exactly how SKN-1 is coupled to a sensor of proteasome activity to induce expression of proteasome subunits was unknown. Lehrbach, conducted a comprehensive genetic screen of C. elegans mutants unable to activate SKN-1 in order to identify the mutated genes.
"The genetic analysis that Nic did was extremely comprehensive," Ruvkun says, "isolating across the genome of C. elegans nearly 100 new mutations that render these animals unable to sense and respond to decreased proteasome activity. These new mutations were mapped by determining the full genome sequence of all of the mutant strains."
Among many mutated genes that were isolated was a protease enzyme called DDI-1, present in many different species, but not previously associated with protein damage signaling. Another essential element of the pathway is a protein called PNG-1, which removes sugar modifications from other proteins. In order for SKN-1 to be activated, DDI-1 must cleave or clip the protein, and PNG-1 must remove a sugar molecule. The mammalian versions of SKN-1 are the transcription factors Nrf1 and Nrf2, and blocking Nrf1 has been proposed as a way to improve protease inhibitor treatment of cancer.
In another paper published simultaneously in eLife, investigators from the University of Tokyo led by Shigeo Murata, MD, PhD, independently discovered that the human version of C. elegans DDI-1, called DDI-2, is needed to couple proteasome deficiency to the upregulation of proteasome genes in human cells. Taken together, these two papers show that the DDI-1/2 protease is essential to sensing proteasome stress in animals from roundworms to humans, making the enzyme an ideal target for drug development.
"We know from the successful development of protease inhibitors to treat HIV and hepatitis C infection that protease enzymes are highly amenable to the development of small-molecule inhibitors," Ruvkun says. "In addition to potentially being powerful enhancers of the proteasome inhibitors being used to treat multiple myeloma, DDI-1/2 inhibitors may be more generally deployed against conditions in which proteasome deregulation has been implicated - for example the variety of diseases caused by aberrant protein aggregation, such as Alzheimer's disease, Parkinson's disease, Huntington's disease, amyotrophic lateral sclerosis and prion diseases."
A professor of Genetics at Harvard Medical School, Ruvkun adds that inactivating mutations in the human version of PNG-1, a protein called NGLY1, have recently been identified as the cause of a rare genetic disease and that the new findings strongly suggest this condition could be treated by reactivation of proteasome genes. The MGH team's discovery that PNG-1 acts with DDI-1 and SKN-1 in the proteasome pathway now places PNG-1/NGLY1 into an increasingly understood genetic pathway. "We have the genetic tools now to ask in C. elegans, how can we cure NGLY1 deficiency?" says Ruvkun. "We are doing these screens right now and are anxiously awaiting what the genome sequence of the mutants will teach us."
More information: Nicolas J Lehrbach et al, Proteasome dysfunction triggers activation of SKN-1A/Nrf1 by the aspartic protease DDI-1, eLife (2016). DOI: 10.7554/eLife.17721