Risk factors for congenital heart defects may lie both inside and outside the heart
Congenital heart defects (CHDs) are a leading cause of birth defect-related deaths. Understanding how genetic alterations cause such defects is complicated by the fact that many of the critical genes are unknown, and those that are known often contribute only small increases in CHD risk.
In new research publishing Sept. 8 in the Open Access journal PLOS Biology, University of California, Irvine biologists Anne Calof and Arthur Lander and colleagues report that the role of genes in CHD is more complex than previously realized and that overall risk is determined by a combination of gene effects both inside and outside of the heart itself.
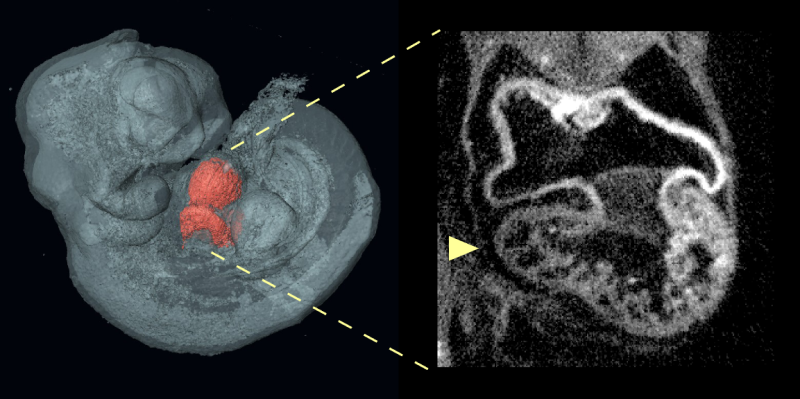
Normal heart formation depends on interactions of multiple types of cells that collaborate in precise times and places throughout development to build the heart's intricate structures. To figure out how these interactions can go awry, the Calof-Lander team studied atrial septal defects (ASDs, a common type of heart defect) in a mouse model of the developmental disorder Cornelia de Lange Syndrome (CdLS).
Most cases of Cornelia de Lange Syndrome are caused by mutations that inactivate a single copy of Nipbl, a gene that directs the expression of many hundreds of other genes in tissues throughout the body. Just as people with Cornelia de Lange Syndrome have a high incidence of heart defects, 30 percent of mice that harbor similar Nipbl mutations exhibit atrial septal defects.
Employing genetically modified mouse models, the researchers used a novel technology to selectively introduce or remove Nipbl mutations in different tissues during embryonic development. Unexpectedly, they found that no Nipbl deficiency in any single tissue—including the tissue that forms the heart itself—could singlehandedly account for the development of atrial septal defects. Rather, the development of heart defects was determined by interactions between heart-forming tissues and the rest of the body. In fact, Nipbl deficiency in some tissues even seemed to protect against the development of atrial septal defects, in certain situations.
In a Primer article that accompanies this research, Bruce Gelb, MD, from the Icahn School of Medicine at Mount Sinai, explains why these were "mind-bending results" and writes that "this work provides novel insights into incomplete penetrance and oligogenic effects underlying CHD." He adds that the novel observations "add further complexity to the way in which we need to think about CHD pathogenesis".
"Our results lead us to hypothesize that heart defects such as ASDs occur when the heart does not grow quickly enough to meet the demands of the developing body - in other words, that heart size and body size must be coordinated for the heart to develop without defects," said Calof, professor of anatomy & neurobiology and developmental & cell biology at UCI. "To our knowledge, this is the first genetic demonstration that major risk factors for heart defects are likely to lie outside of the heart itself."
"When a single gene change causes a birth defect, we often assume that it's because one thing goes wrong in one cell type. The big difference in our studies may have to do with the fact that Nipbl controls a large number of other genes," said Lander, the Donald Bren Professor of Developmental & Cell Biology and director of UCI's Center for Complex Biological Systems. "Given that most human CHDs are now thought to be caused by gene variants acting in combination, what we learned from Nipbl-deficient mice may actually be more typical of the way most CHDs arise."
More information: PLOS Biology, dx.doi.org/10.1371/journal.pbio.2000197