Researchers uncover protein-based 'cancer signature'
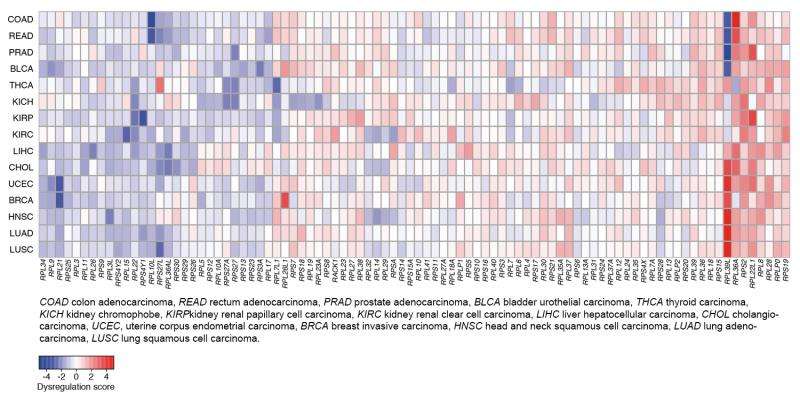
A research team at the University of Basel's Biozentrum has investigated the expression of ribosomal proteins in a wide range of human tissues including tumors and discovered a cancer type specific signature. As the researchers report in Genome Biology, this "cancer signature" could potentially be used to predict the progression of the disease.
Proteins are the building blocks of life. They are produced by molecular machines, called ribosomes. A human ribosome contains some eighty ribosomal proteins. Prof. Mihaela Zavolan's research group at the Biozentrum of the University of Basel has now discovered that about a quarter of the ribosomal proteins have tissue-specific expression and that different cancer types have their own individual expression pattern of ribosomal proteins. In the future, these patterns may serve as a prognostic marker for cancer and may point towards new therapeutic opportunities.
Cellular machines for protein synthesis
Ribosomes are responsible for protein synthesis and are thus essential for the cell. Therefore, it has long been assumed that the expression of the individual components of the ribosomes is strictly controlled and invariant. A few studies, however, have already suggested that the expression of individual ribosomal proteins is altered in cancers as well as in diseases of the hematopoietic system such as acute lymphoblastic leukemia.
"Cancer signature" revealed by systematic data analysis
Mihaela Zavolan and her co-worker Joao Guimaraes have systematically analyzed ribosomal protein expression in thirty tissue types, three hundred different cell types and sixteen different types of tumors, such as lung and breast cancer. In contrast to previous assumptions, they found a wide variability in ribosomal protein gene expression. In particular, hematopoietic and tumor cells display the most complex expression pattern.
"For us, it was really impressive to see that consistent signatures emerged for the different cancer types after the analysis of distinct data sets including patient samples," explains first author Guimaraes. "The pattern of the dysregulated proteins is very striking, whereby the expression of some ribosomal proteins is systematically reduced, and of others increased in cancer cells. This suggests that individual ribosomal proteins can either suppress or promote tumorigenesis."
Expression pattern as a prognostic marker
Furthermore, the scientists discovered a strong relationship between the "signature" in breast cancer and the relapse-free survival. "We were quite surprised to find that the expression level of just three ribosomal proteins allows a fairly accurate prognosis of disease progression, comparable to the best predictive markers that are currently known," Zavolan points out.
"Our study demonstrates the potential of such expression signatures for the prognosis and perhaps a diagnosis of cancer. We are especially interested to study the functions of individual ribosomal proteins and hopefully open the door for new therapeutic options," explains the scientist.
More information: Joao C. Guimaraes et al. Patterns of ribosomal protein expression specify normal and malignant human cells, Genome Biology (2016). DOI: 10.1186/s13059-016-1104-z














