New data network for Huntington's disease research
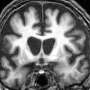
Huntington's disease is an hereditary disorder of the nervous system caused by a faulty gene on chromosome four. The faulty gene leads to cell death in neurons in the brain resulting in gradual physical, mental and emotional changes, and ultimately death. Those born to a parent with Huntington's disease have a 50:50 chance of developing it, and there is currently no cure.
Scientists around the world are working to find effective therapies for Huntington's disease. While the disease is caused by a single gene mutation, the manifestation of the disease at a molecular level is nevertheless highly complex and involves multiple cellular processes. In research, this complexity has resulted in huge and diverse sets of data which in turn pose considerable challenges to researchers in collating, analysing and cross-referencing varied data sources.
An international team of researchers, bioinformaticians and biostatisticians has developed the first freely available data network for scientists researching this debilitating and fatal condition, called HDNetDB. In a paper published in Scientific Reports, the HDNetDB team outlines the challenges facing those investigating Huntington's disease, explains how HDNetDB works and illustrates its effectiveness by discovering potential drug targets, which might help to prevent the neuronal cell death observed in Huntington's disease. A unique feature of this new resource is that it links various cellular processes in a molecular network and provides a more holistic view of the disease.
The team also states that HDNetDB could form the blueprint for similar data networks relating to other specific conditions.
HDNetDB allows users to obtain, visualise and prioritise molecular interaction networks using Huntington's disease-related gene expression and other types of data obtained from human samples and other sources.
Leading the project from the University of Plymouth is Matthias Futschik, Professor in Bioinformatics at the University's School of Biomedical and Healthcare Sciences.
He commented: "Huntington's disease is especially devastating for affected families, since we can nowadays exactly predict who will be affected later in life, but we cannot yet provide any cure. So, it is important that those scientists seeking effective therapies are not hampered by poor access to the vast array of data related to the disease and can connect the different pathological mechanisms. In HDNetDB we have developed a computational tool which allows those scientists free and open access, helping them to identify new molecular targets for the development of new drug strategies for the disease. We believe that our approach can be applied to other neurodegenerative disorders such as Alzheimer's and Parkinson's disease and could facilitate the development of new therapies for these diseases too."
More information: Ravi Kiran Reddy Kalathur et al, HDNetDB: A Molecular Interaction Database for Network-Oriented Investigations into Huntington's Disease, Scientific Reports (2017). DOI: 10.1038/s41598-017-05224-0