Scientists use supercomputer to search for "memory molecules"
Until now, searching for genes related to memory capacity has been comparable to seeking out the proverbial "needle in a haystack." Scientists at the University of Basel made use of the CSCS supercomputer Piz Daint to discover interrelationships in the human genome that might simplify the search for "memory molecules" and eventually lead to more effective medical treatment for people with diseases that are accompanied by memory disturbance.
Every human being's physical and mental constitution is the outcome of a complex interaction between environmental factors and the individual genetic makeup (DNA). The complete set of genes and the genetic information it stores is called the genotype. This genotype influences, among other things, a person's memory and hence the ability to remember the past. In addition to the DNA coding, there are other factors contributing to memory capacity, such as nutrition, schooling and family homelife.
Scientists at the University of Basel from the Transfaculty Research Platform Molecular and Cognitive Neuroscience (MCN) are interested in processes related to memory performance by investigating the molecular basis of memory. "Molecular neuroscience is a dynamic field, with work being done around the world by a community that ranges from mathematicians and computer scientists to applied psychologists," explains Annette Milnik, a post-doctoral fellow in the research group under Professor Andreas Papassotiropoulos, co-head of the research platform. The goal of Milnik's research is to find patterns in genes that are related to memory capacity and that might explain how memory works and how it can be influenced. "There is no such thing as 'the' memory gene, but rather many variations in the genome that, combined with numerous other factors, form our memory," says Milnik.
To investigate memory capacity, researchers from the fields of medical science, psychiatry, psychology and biology make usage of brainwave measurements, memory tests and imaging techniques while the brain is subjected to various stimuli. The researchers also make usage of animal models, as well as genetic and epigenetic studies. The latter examine phenomena and mechanisms that cause chemical changes in the chromosomes and genes without altering their actual DNA sequence.
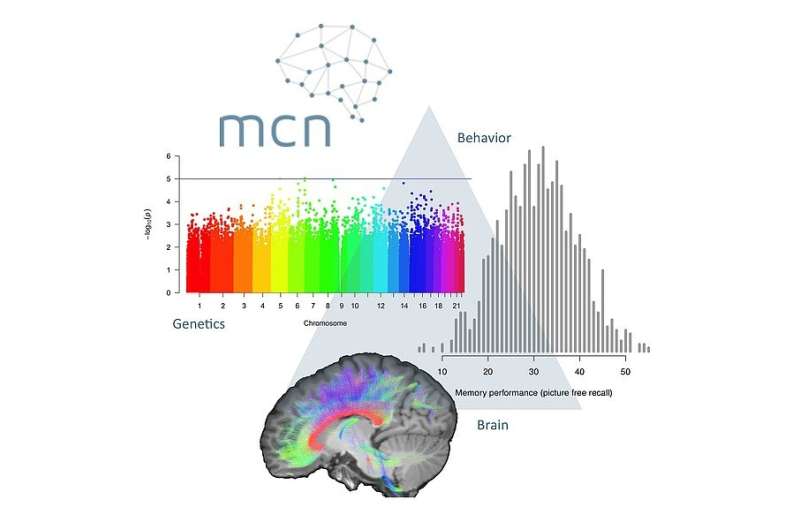
To decipher the molecular basis of memory capacity, researchers "zoom" deep into the human DNA. For this purpose, Milnik examines particular gene segments and their variants. Originally Milnik studied psychology and human medicine, but five years ago she traded in her doctor's career for research and the associated statistical analysis. In her current work, the number of statistical tests together amounted to one quadrillion (1015). Analysing such a quantity of data would not be possible without a supercomputer like "Piz Daint," she notes. Yet her results might significantly simplify future analysis of large datasets in the search for the "memory molecule."
Although the genetic code (DNA) is fixed in all cells, mechanisms like epigenetic processes exists that regulate which parts of the code are expressed. As an example, kidney and liver cells each use different parts of the genome. One process that performs this "functional attribution" is called methylation. "You could imagine flags marking the spots on the human genome where methylation takes place," explains Milnik. A pattern of flags typical for a particular gene thus identifies a given cell function like a pointer. The cell function is influenced by how the genes are specifically read. Moreover, according to Milnik, the environment too may influence the flag-patterns. "We are facing highly complex relationships between genes, the environment, and how they interact. This is why we want to take a step back and seek out a simplified model for these relationships," says Milnik.
Using material sampled from healthy young volunteers, Milnik and her colleagues examined 500,000 genetic variations known as Single Nucleotide Polymorphisms (SNPs)–the basic building blocks of nucleic acids–in conjunction with 400,000 flag-patterns. They wanted to investigate the impact of the genetic code on methylation. According to the researchers, the results of their study show not only that single SNPs located nearby the flags have an impact on the flag-pattern (methylation), but also that combinations of genetic variants–both in proximity and farther apart in the genome–affect this flag-pattern. "This shows us that genetic variants exert a complex influence on methylation," says Milnik. The flag-pattern thus unifies the impact of a larger set of genetic variants that is then represented in one signal.
This means for Milnik that she has found a kind of intermediate filter that reduces the large datasets that have been used to investigate memory capacity. In the past, each single genetic variation has been related individually to memory capacity. But now, according to Milnik, we know that the flags accumulate information from a complex system of SNP-effects. So in the future, instead of using all of the individual SNPs to explore memory capacity and other complex human characteristics, the methylation flag-pattern could be used as well.
This approach as it currently stands is still basic research. However, once the molecules relevant to memory capacity can be identified in this way, a next step could be to investigate whether medicines exist that interact with the corresponding gene products and which might be able to influence memory capacity, explains Milnik. This would offer a gleam of hope for treatment of diseases that are accompanied by memory disturbance, such as dementia or schizophrenia.
More information: Tobias Egli et al. Exhaustive search for epistatic effects on the human methylome, Scientific Reports (2017). DOI: 10.1038/s41598-017-13256-9