New gene expression analysis paves way for improved disease diagnosis and treatment
A comprehensive new analysis from the Max Planck Institute for Biology of Ageing, Germany, and Karolinska Institutet, Sweden, provides insight on how the dysfunction of an important biological process causes disease.
The study in mice, published today in eLife, could open new avenues for research into the deficiency of a system called oxidative phosphorylation (OXPHOS) and provide a valuable reference for future diagnosis and treatment.
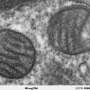
OXPHOS is a metabolic process whereby cells release energy from the food we eat by using enzymes to harvest it and convert it into a molecule called adenosine triphosphate, the cell's fuel. It is produced by structures called mitochondria. Given this central role, mitochondrial dysfunction is a major contributor to human disease and is also involved in the aging process.
"The molecular consequences of OXPHOS dysfunction are hard to predict," says lead author Inge Kühl. "Recent advances in high-throughput technologies in proteomics, metabolomics and sequencing have increased our knowledge of mitochondrial function. However, the events that accompany OXPHOS dysfunction and contribute to mitochondrial diseases are still poorly understood and the treatment options are limited."
To help fill this knowledge gap, Larsson and his team analysed five strains of mice that were deficient in essential factors needed for mitochondrial DNA gene expression in the heart, subsequently leading to OXPHOS dysfunction.
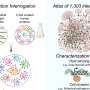
Using an integrated sequencing and mass spectrometry approach, the scientists compared the animals' mitochondrial and cellular proteomes. The team then listed the various gene expression changes at the RNA and protein levels in the mice, caused by OXPHOS dysfunction. Surprisingly, they identified a novel response to OXPHOS dysfunction across all the animals, whereby enzymes in the mitochondria that are necessary for the production of ubiquinone (Q) were severely reduced.
"Q is an essential electron shuttle in the mitochondrial respiratory chain," explains senior author Nils-Göran Larsson, director of the Max Planck Institute for Biology of Ageing and the Institute's Mitochondrial Biology department. "Its deficiency, caused by impaired mitochondrial DNA gene expression in the mouse heart, could potentially be a target for future therapeutics."
"Altogether, our comparative analyses provide a high-quality resource of altered gene expression patterns under OXPHOS deficiency," adds co-author Maria Miranda, PhD student at the Max Planck Institute for Biology of Ageing. "Our datasets can be mined for future studies in this area and will hopefully contribute towards improved patient diagnosis and research on future treatment strategies."
More information: Inge Kühl et al, Transcriptomic and proteomic landscape of mitochondrial dysfunction reveals secondary coenzyme Q deficiency in mammals, eLife (2017). DOI: 10.7554/eLife.30952