Study: Cells of three advanced cancers die with drug-like compounds that reverse chemo failure
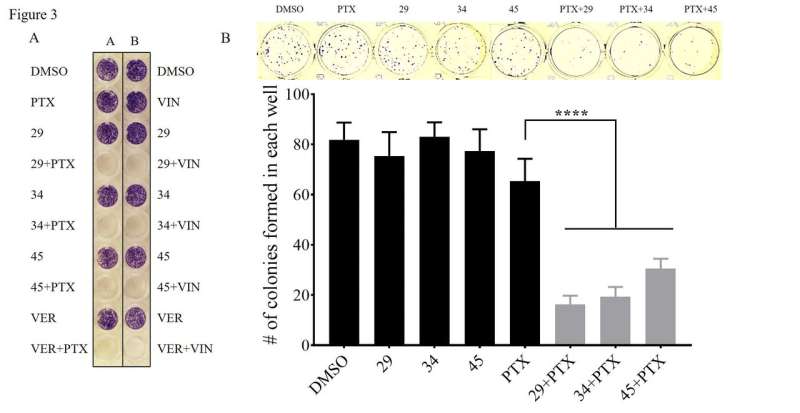
Researchers at Southern Methodist University have discovered three drug-like compounds that successfully reverse chemotherapy failure in three of the most commonly aggressive cancers—ovarian, prostate and breast.
The molecules were first discovered computationally via high-performance supercomputing. Now their effectiveness against specific cancers has been confirmed via wet-lab experiments, said biochemistry professors Pia Vogel and John G. Wise, who led the study.
Wise and Vogel report the advancement in the Nature journal Scientific Reports.
The computational discovery was confirmed in the Wise-Vogel labs at SMU after aggressive micro-tumors cultured in the labs were treated with a solution carrying the molecules in combination with a classic chemotherapy drug. The chemotherapy drug by itself was not effective in treating the drug-resistant cancer.
"Nature designs all cells with survival mechanisms, and cancer cells are no exception," said Vogel, a professor and director of SMU's Center for Drug Discovery, Design and Delivery. "So it was incredibly gratifying that we were able to identify molecules that can inhibit that mechanism in the cancer cells, thereby bolstering the effectiveness of chemotherapeutic drugs. We saw the drugs penetrate these resistant cancer cells and allow chemotherapy to destroy them. While this is far from being a developed drug that will be available on the market anytime soon, this success in the lab gives us hope for developing new drugs to fight cancer."
The current battle to defeat cancer is thwarted by chemotherapy failure in advanced cancers. Cancer cells initially treated with chemotherapy drugs ultimately evolve to resist the drugs. That renders chemotherapy ineffective, allowing cancers to grow and spread.
Key to cancer cell resistance are often certain proteins typically found in all cells—cancerous or otherwise—that are outfitted with beneficial mechanisms that pump away toxins to ensure a cell's continued survival. Nature has set it up that these pumps are prevalent throughout the body, with some areas naturally having more of the pumps than others.
"The cancer cell itself can use all these built-in defenses to protect it from the kinds of things we're using to try to kill it with," Wise said.
The most common of these beneficial defense mechanisms is a pump protein, P-glycoprotein or P-gp, as it's called. Another is one seen in breast and many other cancers, called breast cancer resistance protein, BCRP. In the case of cancer cells on the first round of treatment, these pumps are typically not produced in high levels in the cells, which allows chemotherapy to enter most of the cells in the tumor. This often gives what looks like a good result.
Unfortunately, in the cancer cells that don't die, the chemotherapeutic often changes the cell, which then adapts to protect itself by aggressively multiplying the production of its defensive pumps.
Upon subsequent rounds of chemo, the P-gp and BCRP pumping mechanisms have proliferated. They effectively resist the chemotherapy, which now is much less successful, or not successful at all.
"if enough of the pumps are present, the cancer isn't treatable anymore," said Wise, associate professor in the SMU Department of Biological Sciences. Researchers in the field have searched unsuccessfully for compounds to inhibit the pumps that could be used in the clinic as well.
The molecules that Wise and Vogel discovered stopped the pumps.
"They effectively bring the cancer cells back to a sensitivity as if they'd never seen chemotherapy before," said Vogel. "And our data indicated the molecules aren't cancer specific. They can be used to treat all kinds of cancers because they inhibit not just the P-gp pump, but also the breast cancer protein pump."
To test the compounds, the researchers used amounts of chemotherapeutic that would not kill these multi-drug resistant cancers if the pumps were not blocked.
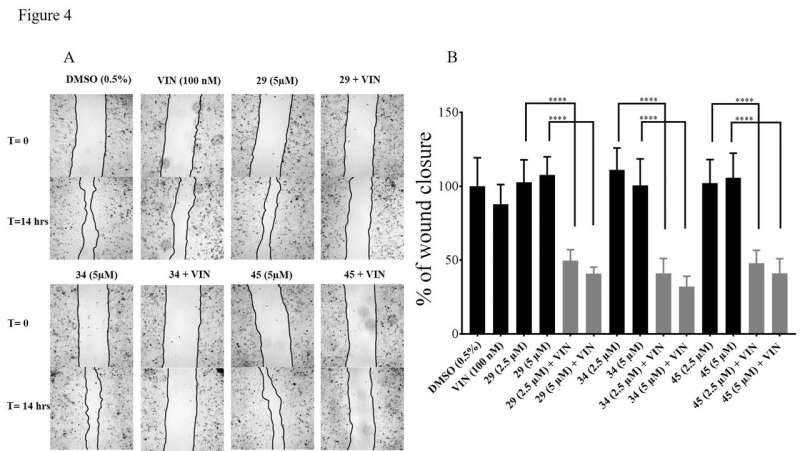
"We wanted to make sure when using these really aggressive cancers that if we do knock out the pump, that the chemotherapy goes in there and causes the cell to die, so it doesn't just stop it temporarily," Wise said. "We spent a fair amount of time proving that point. It turns out that when a cell dies it goes through very predictable morphological changes. The DNA gets chopped up into small pieces, and we can see that, and so the nucleus becomes fragmented, and we can see that. Under the microscope, with proper staining, you can actually see that these highly drug-resistant prostate cancer cells, for example, are dead."
Getting at the heart of the problem
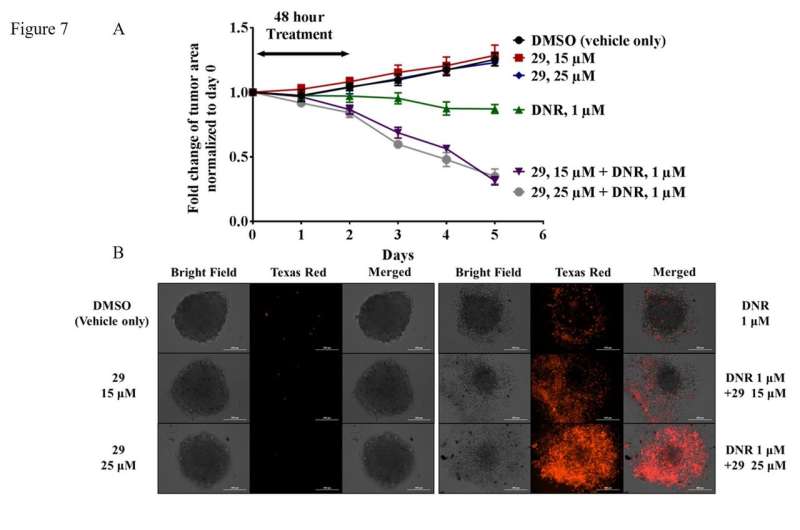
Unique to the experiment is that the molecules were also tested on three-dimensional micro-tumors. That is a departure from the usual cell-culture experiments, which are a two-dimensional film.
In two-dimensional experiments, every cell is exposed to the chemotherapeutic because the film is just one layer of cells thick. That method ignores one of the key challenges to reversing tumors—how to get drugs into the middle of a tumor, not just on its surface.
"We show that with the help of our inhibitor compounds, we actually make the tumor penetrable to chemotherapeutic," Vogel said. "We can kill the cells in the middle of the tumor."
A pathway to personalized medical treatments
Chemotherapy's harmful side effects on non-cancerous organs is well-known. The discovery of molecules that target a specific pump may mitigate that problem.
A patient's tumor can be sampled to see which pump is causing the drug resistance. Then the molecule that knocks out that specific pump can be added to the chemotherapy.
"That means you don't open the door wide to toxins in the central nervous system," Wise said. "That has some real implications for the future and for personalized medicine. In most of the previous clinical trials, inhibitors have opened the brain up to toxins. From what we can tell so far, our inhibitors do not increase the toxicity of chemotherapeutics in normal cells."
An audacious discovery
P-gp is present in one form or another in everything that lives.
"It's in your dog, it's in your cat, it's in yeast cells, it's in bacteria, it's everywhere," Wise said. "Why is it everywhere? Because it's a really wonderful solution to the problem of getting toxins out of a cell. P-gp is a tremendously sophisticated evolutionary solution to that problem. And as with most things in biology that work well, everybody gets it, because if you don't have it, you didn't survive."
Biologists say that P-gp can pump out 95 of 100 chemotherapeutics, indicating it can grab almost any drug and throw it out of a cell.
"So there's a certain audacity to say that we can use a computer and target one part of this protein—the motor—and totally avoid the part of the protein that has evolved to pump almost anything that looks like a drug out of the cell," Wise said. "That's an audacious claim and the findings surprised us."
In their computational and wet-lab experiments, Wise and Vogel searched for molecules that inhibit ATP hydrolysis—the chemical energy reaction that powers the P-gp pump.
"We targeted the motor of the pump instead of the pump part of the pump because almost all the clinical trial failures in other studies were actually compounds that targeted the pump part of the pump—and they would just slow down the pumping of the chemotherapeutic," Vogel said. "The time was ripe to do these structural models. We hypothesized that we could completely avoid the pumping mechanism and just target the motor."
Computational method highly predictive
The wet-lab experiments confirmed the accuracy of the computational findings, Vogel said.
"The predictiveness of the computational methods was really high," she said. "It completely exceeded my expectations. We had selected certain molecules that were predicted in those computational experiments to interact with the pump in certain ways and not in others, and we could show in our wet-lab experiments that the predictions were spot on."
Fascinated by the novel approach to the research, the National Institute of General Medical Sciences funded much of the research.
Wise and Vogel tapped the high-performance computing power of SMU's ManeFrame, one of the most powerful academic supercomputers in the nation. Wise sorted through 15 million commercially available drug-like compounds made publically available in digital form from the pharmacology database Zinc at the University of California, San Francisco.
Then, again using ManeFrame, Wise ran the compounds through a computer-generated model of P-gp. The virtual model, designed and built by Wise, is the first computational microscope of its kind to simulate the actual behavior of P-gp in the human body, including interactions with drug-like compounds while taking on different shapes. He reported the dynamic functioning of the model in 2015 in the journal Biochemistry.
Process of elimination finds needle in the haystack
Out of 15 million drug-like compounds that were virtually screened, the researchers found 180,000 that in the computer were predicted to interact strongly with the ATP harvesting power plant part of the pump motor. From those, Wise and Vogel eliminated the ones that interact well with the pump part. Roughly 0.15 percent survived—several hundred.
"So that tells you how promiscuous that binding site is for compounds," Wise said.
From there, they bought and tested in the lab a number of the remaining molecules.
"It was a process of elimination," Vogel said. "Of the first 38 we tested, we found four. And because of the computational approach we took, it made failure relatively cheap. This is proof of principle that at least in those cases the compounds behave exactly in the lab as predicted in the computer. Which thrills the heck out of me—I never, ever would have thought that."
The Vogel and Wise research labs are part of the Center for Drug Discovery, Design and Delivery in SMU's Dedman College. The center's mission is a novel multi-disciplinary focus for scientific research targeting medically important problems in human health.
More information: Amila K. Nanayakkara et al, Targeted inhibitors of P-glycoprotein increase chemotherapeutic-induced mortality of multidrug resistant tumor cells, Scientific Reports (2018). DOI: 10.1038/s41598-018-19325-x