ID'ing features of flu virus genome may help target surveillance for pandemic flu
The current influenza outbreak—the worst across the United States in nearly a decade—is worrisome but still far less dire than a pandemic flu, which could kill millions. Such pandemics are exceedingly difficult to predict, but new research at Washington University School of Medicine in St. Louis offers details about flu viruses that could help improve surveillance to detect a potential pandemic.
Pandemic flu occurs when flu strains from different species—birds and humans, or humans and pigs—genetically mix to make a new virus that spreads faster and makes people sicker than either strain alone. Public health authorities monitor places where people live in close contact with animals for the first signs of new pandemic viruses.
Reporting Jan. 31 in Nature Communications, the researchers identified features of the influenza virus genome that affect how well the virus multiplies. These features are similar but not identical across viral strains. It's possible that the extent of similarity between strains influences whether two flu viruses can mix their genetic material to make a hybrid virus with the potential to explode into pandemic flu.
"We think that two strains need to have similar features in their genome to re-assort and make a new virus," said senior author Jacco Boon, Ph.D., an assistant professor of medicine at Washington University. "We hope that in the future, this work will allow us to focus on certain strains of influenza virus and target our surveillance more narrowly so we have a better chance of identifying the next pandemic flu before it spreads."
Flu viruses multiply by infecting cells and hijacking the cell's machinery to mass-produce copies of the virus's genome and proteins, which are then bundled into new viruses. Influenza virus's genome is broken into eight parcels of RNA, a molecule similar to DNA. When a cell is infected with two or more flu strains at once, the genetic parcels from the different strains tend to get mixed up. The result is often a new influenza strain born with genetic information from multiple parental strains.
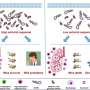
Boon and first author Graham Williams, Ph.D., now a postdoctoral researcher at Duke University, with the help of Sebla Kutluay, Ph.D., an assistant professor of molecular microbiology, found that parts of the virus's RNA genome fold like origami into specific 3-D shapes and that these shapes are necessary for the virus to multiply. When they mutated the genome to change the shapes, the viruses did not reproduce well. "Silent" mutations that left the shapes intact, on the other hand, did not affect multiplication.
There are thousands of different flu viruses in the world, each differing slightly in their genetic sequence and, most likely, the shapes into which their RNA folds. Flu viruses whose genomes form very different 3-D structures may not be able to recombine into a new strain.
"Right now we do surveillance on pretty much everything," said Boon, who is also an assistant professor of molecular microbiology, and of pathology and immunology. "But if we know that the viruses from a certain species or a certain region just don't have the right RNA features, then we can make surveilling them a lower priority. If we can focus our resources more effectively, we may be able to catch the next pandemic flu before it really gets going."
More information: Graham D. Williams et al, Nucleotide resolution mapping of influenza A virus nucleoprotein-RNA interactions reveals RNA features required for replication, Nature Communications (2018). DOI: 10.1038/s41467-018-02886-w