PARP-1 may be key to effectiveness of PARP inhibitors, and now researchers can image it
Penn Medicine researchers have used CRISPR/Cas9 gene editing technology to isolate a key genetic feature that could cause resistance to PARP inhibitors in patients with ovarian cancer—and they've also proven they have a way to see that feature using PET imaging.
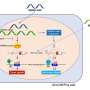
The team found PARP inhibitors—a type of targeted therapy that kills cancer cells with mutations in their DNA repair genes while sparing healthy tissue that does not have the mutations—specifically require the presence of PARP-1 in order to take effect. They also show that a radioactive tracer developed at Penn makes PARP-1 visible during PET scans and may provide a method of measuring PARP-1 in ovarian cancer that complements biopsy. The findings of this multidisciplinary team—which included radiologists, pathologists, and oncologists from the University of Pennsylvania's Perelman School of Medicine—were published in the Journal of Clinical Investigation this month.
Epithelial ovarian cancer is the fifth deadliest type of cancer in women, with more than 70 percent of patients already showing advanced disease when they're diagnosed. Standard treatment relies on chemotherapy, but more recently, PARP inhibitors have emerged as a way to improve patient outcomes. Three of these drugs are currently approved for use by the U.S. Food and Drug Administration, and patients commonly receive these inhibitors if they test positive for the genetic mutations BRCA1 and BRCA2. However, only about 50 percent of these patients respond.
Penn researchers used CRISPR/Cas9 gene editing in the lab to delete PARP-1 from ovarian cancer cells, then exposed those cells to PARP inhibitors and compared them to a control group with PARP-1. Cells without PARP-1 showed less DNA damage and less responsiveness to the inhibitors than the unedited control group cells. In one case, the loss of PARP-1 resulted in a greater than a 1,000-fold decrease in sensitivity to the PARP inhibitor.
"These findings show that PARP-1 is required for PARP inhibitors to do their job," said the study's co-senior author Mehran Makvandi, PharmD, RPh, BCNP, an instructor of Radiology. "These data also demonstrated the clinical need to evaluate PARP-1 levels in BRCA cancers to better understand which patients are likely to benefit and which may be resistant."
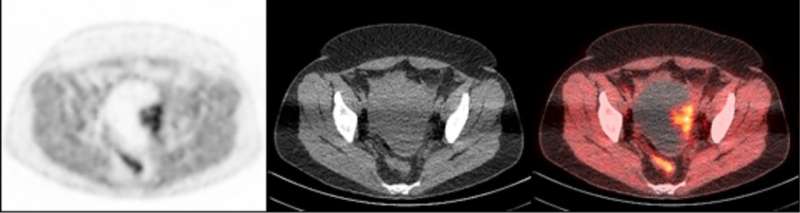
To accomplish this, researchers used a radioactive tracer that binds to PARP-1, making it visible during PET scans. The tracer, called FluorThanatrace, was specially designed by Robert H. Mach, Ph.D., the Britton Chance Professor of Radiology. After a pre-clinical test using mouse models showed the tracer was hitting its specific molecular target, the study's authors moved to a phase I trial.
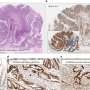
A clinical trial involving 10 patients led to 13 tissue samples. These samples had a wide spectrum of uptake of the tracer, but that uptake correlated with PARP-1 expression. In other words, the more the tracer was able to bind to the tumor, the higher the level of PARP-1 that tumor expressed.
"This is proof-of-concept that we not only have a potential biomarker for the effectiveness of PARP inhibitors, but we also have a non-invasive imaging technique to find that biomarker in ovarian cancer patients," Makvandi said.
Researchers say the next step in this work is to evaluate whether this technique can be used to predict efficacy and resistance to PARP inhibitors. Makvandi and Fiona Simpkins, MD, an assistant professor of Obstetrics and Gynecology and co-author on this study, are leading future clinical trials to directly test the predictive value of this approach in ovarian cancer patients receiving PARP inhibitor monotherapy. Trials are also planned in breast cancer.
More information: Mehran Makvandi et al, A PET imaging agent for evaluating PARP-1 expression in ovarian cancer, Journal of Clinical Investigation (2018). DOI: 10.1172/JCI97992