Therapeutic RNA corrects splicing defect that causes familial dysautonomia
Scientists at Cold Spring Harbor Laboratory (CSHL) have identified a therapeutic RNA molecule that corrects the error in genetic processing that leads to familial dysautonomia, a rare inherited neurodegenerative disorder. The experiments, conducted in cells sampled from patients and in a mouse model of the disease, provide proof of the team's therapeutic concept.
Familial dysautonomia, or FD, occurs primarily in people of Ashkenazi Jewish ancestry. It impairs the development and function of the autonomic nervous system, causing problems with involuntary body functions, such as breathing, digestion, and blood-pressure regulation.
Those affected carry a mutation in a gene called IKBKAP, which reduces cells' ability to produce the IKAP protein. Now, researchers led by CSHL Professor Adrian Krainer, together with their collaborators at Ionis Pharmaceuticals, have used a synthetic piece of RNA to enable human and mouse cells to produce normal levels of IKAP, despite the presence of the disease mutation.
The RNA-like molecule—known as an antisense oligonucleotide (ASO)—was injected into mice whose cells carry the human IKBKAP mutation that causes FD. It corrects an error in what scientists call RNA splicing that prevents people with FD from manufacturing sufficient functional IKAP protein. A drug based on the same ASO might be used to overcome the same mutation in people with familial dysautonomia, Krainer says.
The findings, reported today in Nucleic Acids Research, come less than two years after the FDA approved nusinersen (marketed as Spinraza), an ASO that Krainer's lab developed in collaboration with Ionis Pharmaceuticals for the treatment of another neurodegenerative disorder called spinal muscular atrophy (SMA).
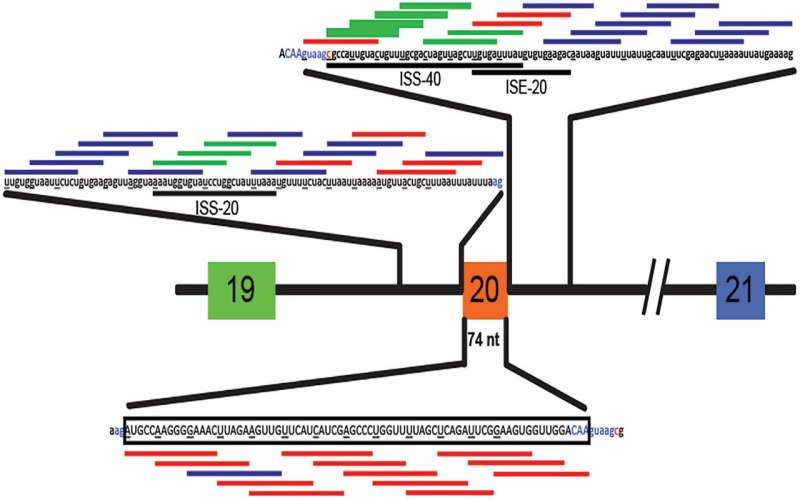
In their work on SMA, Krainer's team had shown that an ASO designed to bind to an RNA near a defective splice site could correct splicing and induce production of functional, full-length protein. They followed the same strategy in the newly published research on FD.
They designed and screened dozens of ASOs that bind to IKBKAP's RNA, looking for one that most efficiently restored normal splicing. "We used the same chemistry as we did for Spinraza, because we know that drug is well tolerated and it's very effective and stable in the central nervous system," Krainer says.
The team demonstrated that when they administered the ASO to mice harboring the mutant human IKBKAP gene, it reached various tissues, including the nervous system, and corrected splicing. More importantly, it restored production of full-length IKAP protein.
Further testing is needed to determine whether it will be appropriate to move the ASO into clinical trials. But following the success of nusinersen, Krainer is optimistic that his team has identified a promising strategy for potentially treating familial dysautonomia.
More information: Sinha R et al, "Antisense oligonucleotides correct the familial dysautonomia splicing defect in IKBKAP transgenic mice" appeared online in Nucleic Acids Research on April 17, 2018.