Researchers describe a new regulatory mechanism for salmonella virulence with small RNA fragments
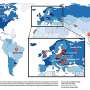
A series of small RNA fragments are the regulator factors involved in a new control mechanism for salmonella virulence, a pathogenic bacterium which causes bacterial gastroenteritis with a high incidence –more than 100,000 cases per year- in Europe.
The findings of the new regulating mechanism, published in the journal PLOS Genetics, are reported by Carlos Balsalobre, lecturer at the Faculty of Biology of the University of Barcelona, and it means a step forward in the development of new therapeutic strategies to treat infections in the food chain.
According to the study, these small RNA chains could regulate the genic expression of the involved genes in a key process in salmonella's ability to cause an infection. Other participants of the study are the UB experts Youssef El Mouali, Tania Gaviria Cantín and Marta Gilabert, as well as María Antonia Sánchez-Romero (University of Seville), and Alexander J. Westermann and Jörg Vogel (University of Würzburg, Germany).
Salmonella enterica is one of the main enteric pathogens in developed countries and underdevelopment areas as well. In fact, salmonella is present in domestic and wild animals, and can cross the whole food chain. Regarding humans, most of the cases of salmonella have their origins in the consumption of foods with natural origin that are already infected by the bacterium, and can cause light gastroenteritis and more general severe infections.
During the infection process, S. enterica invades the epithelial cells, a determining process to unchain the bacterial infection, which requires the coordinated expression of a series of genes. According to Carlos Balsalobre, lecturer at the Department of Genetics, Microbiology and Statistics of the UB, "once the we eat the food that is already affected by salmonella, the bacterium gets to the intestine, where it is able to invade epithelial cells, a determining process to unchain the bacterial infection".
Objective: genes from the Salmonella pathogenicity island
Most of the coded genes by involved factors in the invasion of epithelial cells –and determining for the virulent capacity of the bacterium- are grouped in a chromosomal region known as the Salmonella pathogenicity island (SPI-1). For decades, great part of the scientific research has focused on the description of mechanisms that activate the expression of the genes in SPI-1 during the bacterial infectious process.
"However, in this study we focused on the opposite case", adds Balsalobre. "That is, we studied the important factors to keep these genes silent when the bacterium does not need to invade the guest's epithelial cells".
The new study reveals that the CRP-AMPc metabolic sensor is involved in the control of the expression of the genes in the SPI-1. This control occurs at a post-transcriptional level and indirectly, through the regulation of the expression of a small RNA fragment (Spot 42). The mechanism of the regulation, described now for the first time, takes place through the interaction of the small RNA fragments with the terminal region of the corresponding RNA messenger (in particular, with 3'UTR).
This study, a new step towards the knowledge of control mechanisms in bacteria to regulate the genic expression, will contribute to design new therapeutic tools to work on one of the microorganisms that shows a higher resistance to the use of antimicrobials.
More information: Youssef El Mouali et al. CRP-cAMP mediates silencing of Salmonella virulence at the post-transcriptional level, PLOS Genetics (2018). DOI: 10.1371/journal.pgen.1007401